Probe CUST_37784_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_37784_PI426222305 | JHI_St_60k_v1 | DMT400077052 | CTTGTTTACGCCTCGTTATAATGTTCTTATGGAAAATCACAAAGAGACATGAGTTTGAGT |
All Microarray Probes Designed to Gene DMG400029966
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_37784_PI426222305 | JHI_St_60k_v1 | DMT400077052 | CTTGTTTACGCCTCGTTATAATGTTCTTATGGAAAATCACAAAGAGACATGAGTTTGAGT |
| CUST_37778_PI426222305 | JHI_St_60k_v1 | DMT400077053 | CCTCCATTTGAACTATTGTGTACTTGACTAGCACATGTACTGCCTAGTATTGTTAGTGTC |
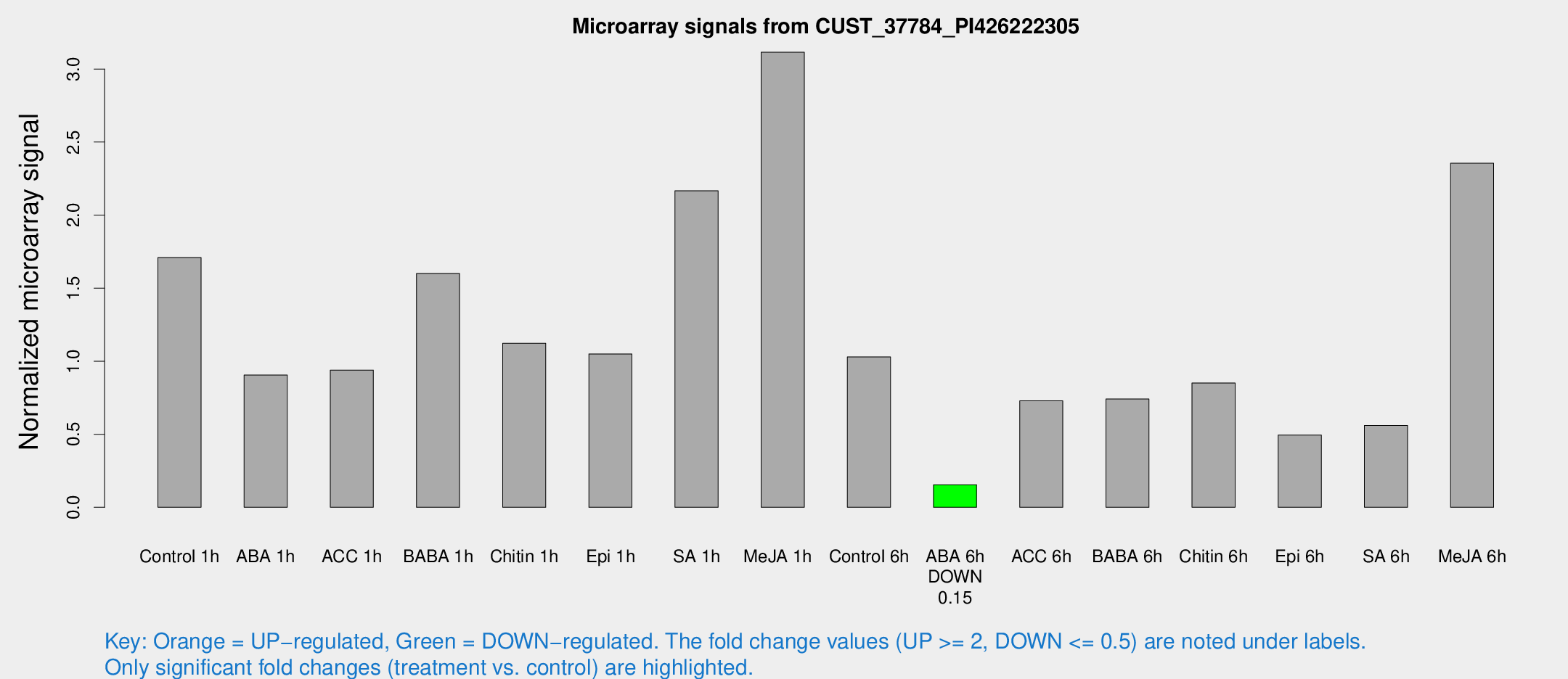
Microarray Signals from CUST_37784_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 92.9041 | 24.7809 | 1.70935 | 0.402992 |
| ABA 1h | 42.5948 | 8.64569 | 0.905178 | 0.151682 |
| ACC 1h | 55.8982 | 20.9262 | 0.938952 | 0.317329 |
| BABA 1h | 85.8762 | 22.6652 | 1.60072 | 0.365472 |
| Chitin 1h | 53.7656 | 12.3951 | 1.12221 | 0.24801 |
| Epi 1h | 46.6885 | 5.63379 | 1.04961 | 0.0964356 |
| SA 1h | 124.535 | 40.7046 | 2.16625 | 0.761289 |
| Me-JA 1h | 147.12 | 47.725 | 3.11505 | 1.12586 |
| Control 6h | 57.3224 | 17.7908 | 1.02906 | 0.267158 |
| ABA 6h | 8.49707 | 3.59591 | 0.154421 | 0.069001 |
| ACC 6h | 42.5763 | 4.82963 | 0.728573 | 0.0933372 |
| BABA 6h | 43.9653 | 9.72804 | 0.740922 | 0.189213 |
| Chitin 6h | 55.163 | 21.5146 | 0.851133 | 0.425427 |
| Epi 6h | 29.5211 | 5.91892 | 0.493862 | 0.0847258 |
| SA 6h | 41.5214 | 18.0866 | 0.559798 | 0.475629 |
| Me-JA 6h | 121.056 | 16.2721 | 2.35497 | 0.444385 |
Source Transcript PGSC0003DMT400077052 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G10310.1 | +2 | 7e-125 | 381 | 245/495 (49%) | high-affinity K+ transporter 1 | chr4:6392008-6395667 FORWARD LENGTH=506 |
