Probe CUST_37612_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_37612_PI426222305 | JHI_St_60k_v1 | DMT400049515 | CCGAATGGGAGTCTTGATTCATTTATATTTGGTACTATTCTCCATCCAATATTGTGCAAA |
All Microarray Probes Designed to Gene DMG401019239
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_37612_PI426222305 | JHI_St_60k_v1 | DMT400049515 | CCGAATGGGAGTCTTGATTCATTTATATTTGGTACTATTCTCCATCCAATATTGTGCAAA |
| CUST_37634_PI426222305 | JHI_St_60k_v1 | DMT400049512 | TACATAGAAGGAAATGCACCGGGTGGTGATTTCACTCCTAGCCAATCTGCACAGATTACA |
Microarray Signals from CUST_37612_PI426222305
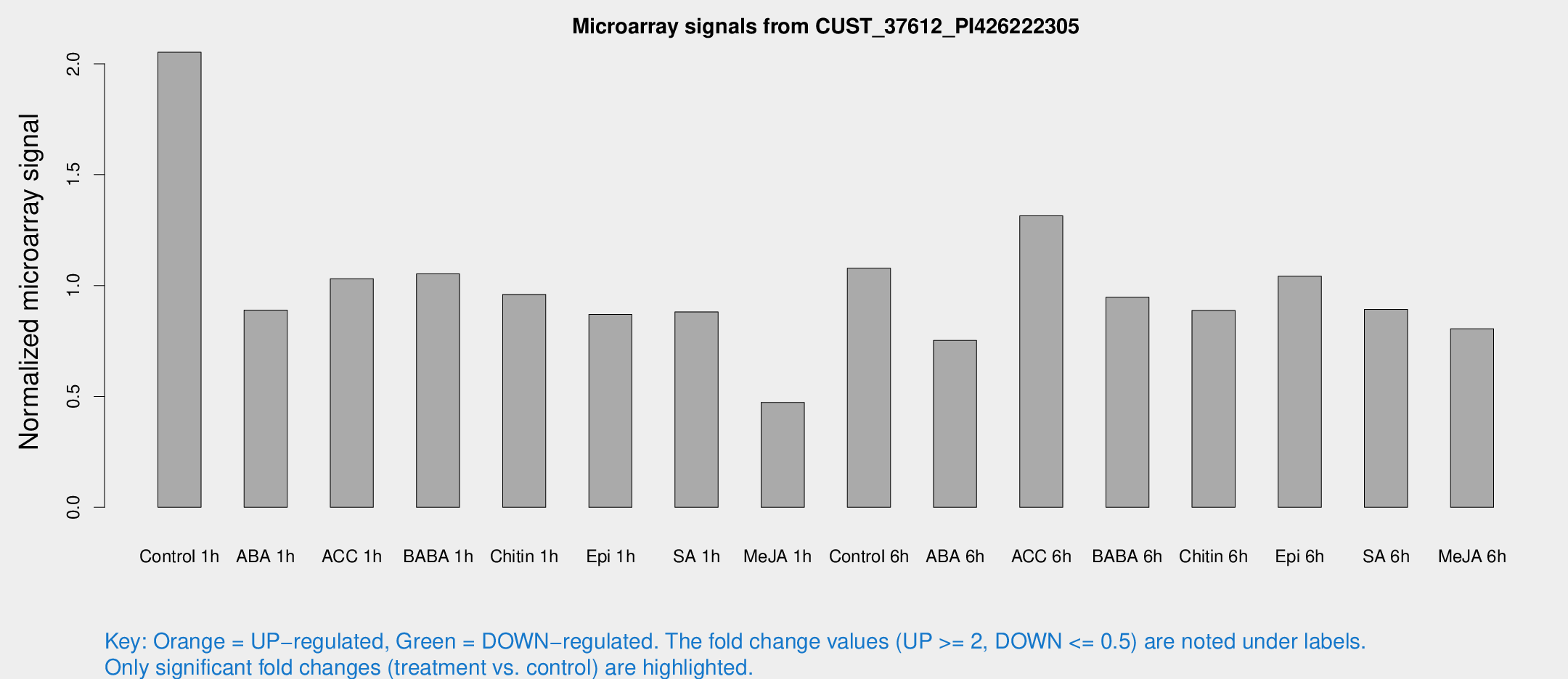
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 45.8071 | 10.1078 | 2.05214 | 0.590071 |
| ABA 1h | 22.3348 | 10.7048 | 0.88907 | 0.629636 |
| ACC 1h | 26.6577 | 9.10412 | 1.0308 | 0.455272 |
| BABA 1h | 25.9935 | 9.20684 | 1.0527 | 0.458633 |
| Chitin 1h | 19.7076 | 5.94884 | 0.959879 | 0.220699 |
| Epi 1h | 16.7349 | 3.59172 | 0.869586 | 0.213844 |
| SA 1h | 20.1226 | 4.28907 | 0.881083 | 0.17987 |
| Me-JA 1h | 8.96921 | 3.44269 | 0.473311 | 0.2144 |
| Control 6h | 23.3617 | 4.08785 | 1.07773 | 0.182585 |
| ABA 6h | 17.5909 | 3.70921 | 0.752556 | 0.174292 |
| ACC 6h | 36.685 | 14.7455 | 1.31455 | 0.678095 |
| BABA 6h | 24.6711 | 6.87131 | 0.94746 | 0.30921 |
| Chitin 6h | 20.8081 | 4.71969 | 0.887384 | 0.191767 |
| Epi 6h | 25.7482 | 4.91927 | 1.04177 | 0.186682 |
| SA 6h | 18.9669 | 3.79314 | 0.892632 | 0.238361 |
| Me-JA 6h | 21.5802 | 8.11239 | 0.805011 | 0.470326 |
Source Transcript PGSC0003DMT400049515 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G27290.1 | +3 | 1e-143 | 441 | 211/416 (51%) | S-locus lectin protein kinase family protein | chr4:13666281-13669202 FORWARD LENGTH=783 |
