Probe CUST_37587_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_37587_PI426222305 | JHI_St_60k_v1 | DMT400049621 | AGACAATACAAGCCGTGTTGTTGGAACATATGGTTACATGTCCCCGGAATATGCAGTCGA |
All Microarray Probes Designed to Gene DMG400019276
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_37587_PI426222305 | JHI_St_60k_v1 | DMT400049621 | AGACAATACAAGCCGTGTTGTTGGAACATATGGTTACATGTCCCCGGAATATGCAGTCGA |
| CUST_37657_PI426222305 | JHI_St_60k_v1 | DMT400049622 | CATCCTTGTCGATTGTGAGTGTCATTCATTTCTCAGAATTGTATTAAAATCCTCCCTTTT |
Microarray Signals from CUST_37587_PI426222305
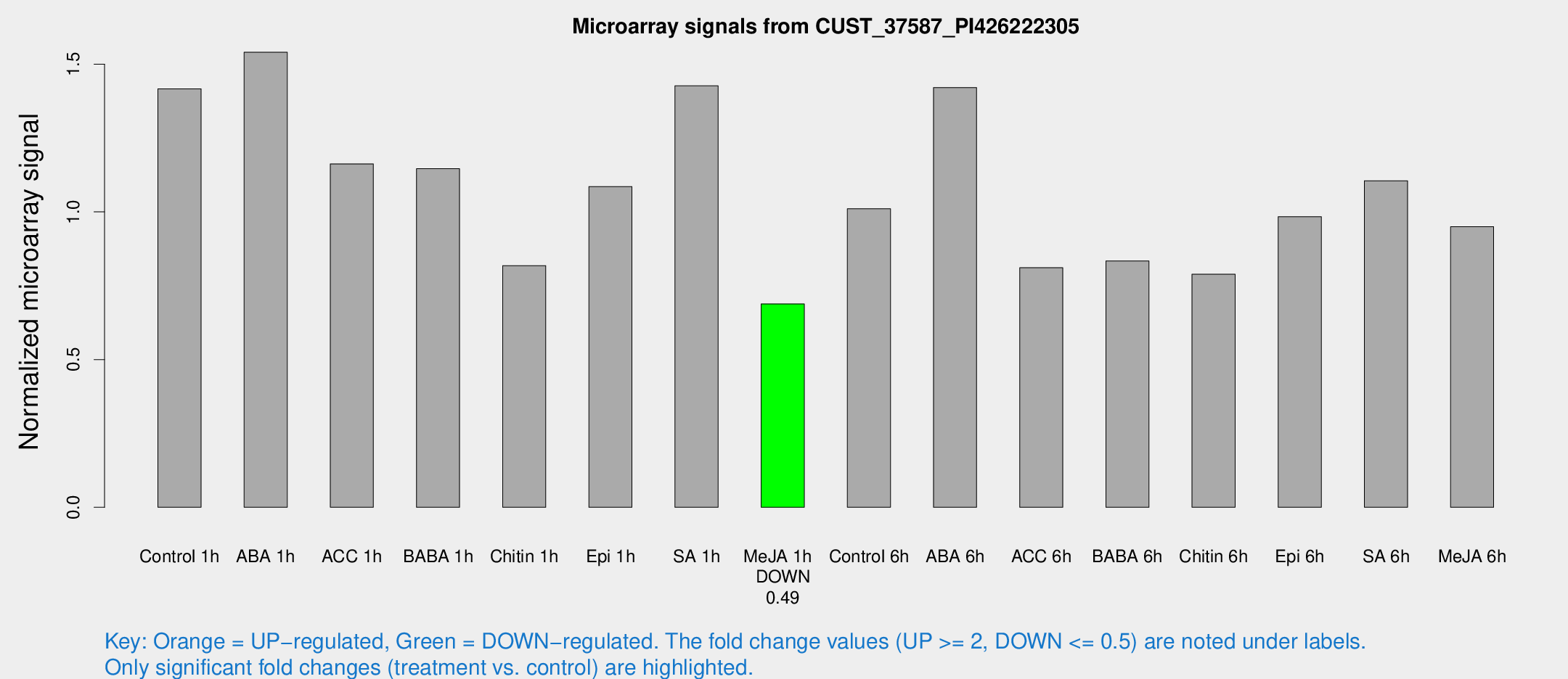
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 656.61 | 106.325 | 1.41665 | 0.13621 |
| ABA 1h | 624.426 | 69.5482 | 1.54034 | 0.0892773 |
| ACC 1h | 597.369 | 162.58 | 1.16248 | 0.321214 |
| BABA 1h | 532.537 | 124.819 | 1.14599 | 0.188834 |
| Chitin 1h | 341.477 | 56.8995 | 0.818004 | 0.0718486 |
| Epi 1h | 425.637 | 24.8041 | 1.08579 | 0.0632088 |
| SA 1h | 678.563 | 98.7329 | 1.42676 | 0.137891 |
| Me-JA 1h | 259.995 | 39.7277 | 0.688495 | 0.0451931 |
| Control 6h | 507.798 | 149.634 | 1.01067 | 0.268486 |
| ABA 6h | 716.476 | 141.523 | 1.42083 | 0.267105 |
| ACC 6h | 421.234 | 24.6737 | 0.811019 | 0.095616 |
| BABA 6h | 429.622 | 56.5467 | 0.833953 | 0.0929832 |
| Chitin 6h | 381.327 | 23.4591 | 0.788862 | 0.0461924 |
| Epi 6h | 524.296 | 106.413 | 0.983668 | 0.171719 |
| SA 6h | 529.502 | 142.587 | 1.10543 | 0.211092 |
| Me-JA 6h | 443.23 | 82.0807 | 0.949947 | 0.0992728 |
Source Transcript PGSC0003DMT400049621 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G27300.1 | +1 | 1e-141 | 439 | 218/342 (64%) | S-locus lectin protein kinase family protein | chr4:13669308-13672348 REVERSE LENGTH=815 |
