Probe CUST_37475_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_37475_PI426222305 | JHI_St_60k_v1 | DMT400078988 | TTTTTGTTGGCGCACATTAGCAGAACGTCATTATTCTTGCTAACATGCGCACCCTACCCA |
All Microarray Probes Designed to Gene DMG402030736
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_37475_PI426222305 | JHI_St_60k_v1 | DMT400078988 | TTTTTGTTGGCGCACATTAGCAGAACGTCATTATTCTTGCTAACATGCGCACCCTACCCA |
| CUST_37514_PI426222305 | JHI_St_60k_v1 | DMT400078987 | TCACAAAATAAGAGGAGTGCCAAGGGATTCAAAAGTCTAGGGCAATTGAGGGGCCTGATT |
Microarray Signals from CUST_37475_PI426222305
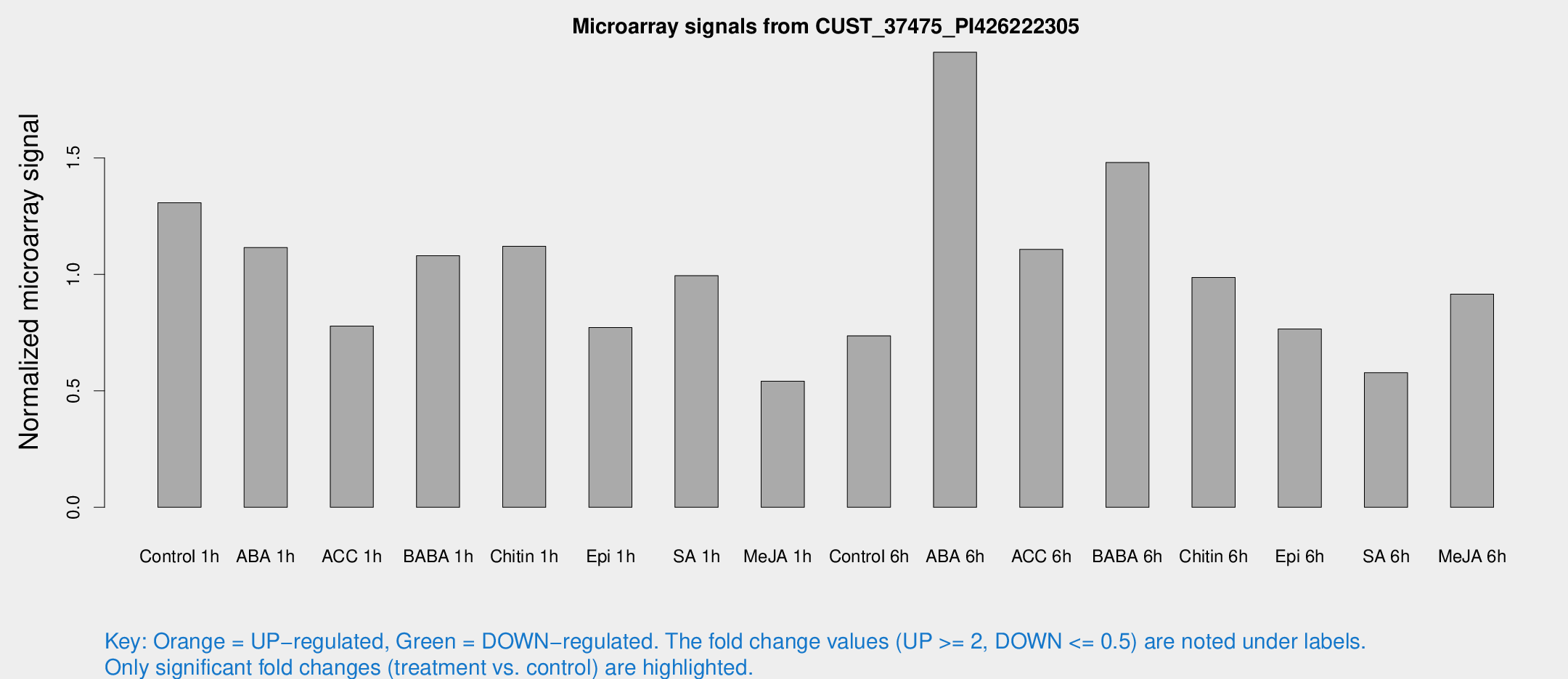
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 31.985 | 9.08697 | 1.30763 | 0.298814 |
| ABA 1h | 22.8316 | 3.33061 | 1.11549 | 0.170634 |
| ACC 1h | 21.2063 | 7.51379 | 0.778074 | 0.288931 |
| BABA 1h | 29.1815 | 10.7349 | 1.08059 | 0.493853 |
| Chitin 1h | 23.7387 | 4.27447 | 1.12081 | 0.224546 |
| Epi 1h | 15.175 | 3.32109 | 0.772229 | 0.168935 |
| SA 1h | 23.9449 | 4.01567 | 0.994603 | 0.1554 |
| Me-JA 1h | 10.4498 | 3.38984 | 0.541163 | 0.188566 |
| Control 6h | 20.0628 | 6.85419 | 0.736429 | 0.304647 |
| ABA 6h | 53.498 | 15.8313 | 1.95418 | 0.725919 |
| ACC 6h | 29.8905 | 5.28124 | 1.10763 | 0.399357 |
| BABA 6h | 38.3622 | 5.2073 | 1.48034 | 0.172861 |
| Chitin 6h | 24.0556 | 4.04544 | 0.986833 | 0.170168 |
| Epi 6h | 21.6017 | 6.66854 | 0.765658 | 0.225743 |
| SA 6h | 19.6054 | 12.5781 | 0.577846 | 0.441953 |
| Me-JA 6h | 21.2456 | 3.57624 | 0.91543 | 0.161928 |
Source Transcript PGSC0003DMT400078988 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G15580.1 | +3 | 8e-10 | 58 | 42/94 (45%) | RING/U-box superfamily protein | chr2:6797687-6798815 FORWARD LENGTH=196 |
