Probe CUST_37450_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_37450_PI426222305 | JHI_St_60k_v1 | DMT400078874 | ACTTTCATAGCAAAGATCTGGCTTGTCTGTCTTGCCTGCAATTAAACTTCATTGTTTTTT |
All Microarray Probes Designed to Gene DMG400030695
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_37444_PI426222305 | JHI_St_60k_v1 | DMT400078873 | AAACCGGAGGAAGTAGATACTAGCAACAATGTTTTACATGTATGTCCACTTTGCAGAGGA |
| CUST_37450_PI426222305 | JHI_St_60k_v1 | DMT400078874 | ACTTTCATAGCAAAGATCTGGCTTGTCTGTCTTGCCTGCAATTAAACTTCATTGTTTTTT |
Microarray Signals from CUST_37450_PI426222305
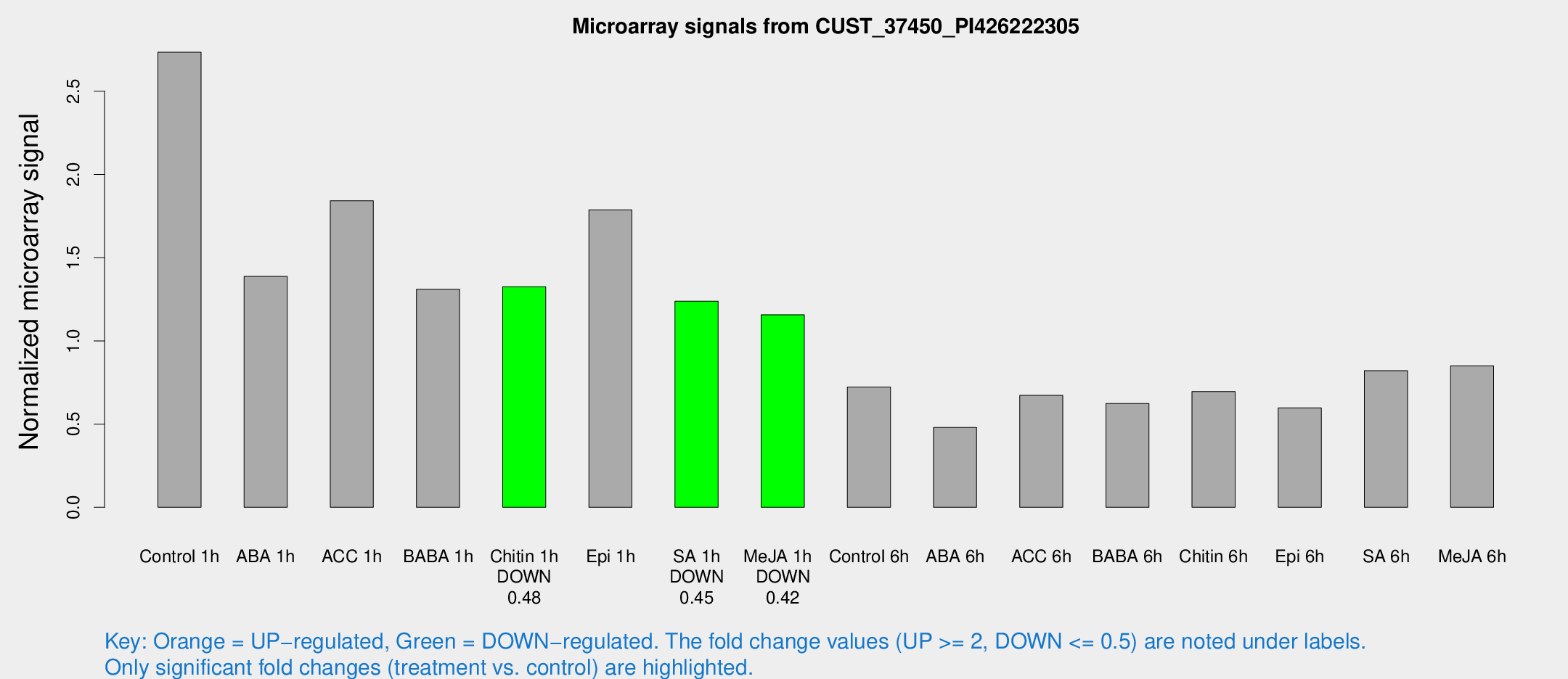
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 207.782 | 14.4302 | 2.73466 | 0.163602 |
| ABA 1h | 94.5378 | 13.1791 | 1.38754 | 0.201681 |
| ACC 1h | 144.938 | 16.5977 | 1.842 | 0.137089 |
| BABA 1h | 105.596 | 29.3052 | 1.3106 | 0.31633 |
| Chitin 1h | 90.3124 | 6.11255 | 1.32606 | 0.134252 |
| Epi 1h | 119.124 | 16.5475 | 1.78741 | 0.258729 |
| SA 1h | 96.7456 | 6.77846 | 1.23889 | 0.0823485 |
| Me-JA 1h | 72.7986 | 9.52308 | 1.15705 | 0.108392 |
| Control 6h | 58.3241 | 15.2968 | 0.722842 | 0.162743 |
| ABA 6h | 38.5539 | 4.13315 | 0.479333 | 0.0514961 |
| ACC 6h | 62.3172 | 14.1803 | 0.672602 | 0.150134 |
| BABA 6h | 52.8443 | 4.77538 | 0.62382 | 0.0562705 |
| Chitin 6h | 56.378 | 5.00096 | 0.695668 | 0.0621211 |
| Epi 6h | 51.002 | 4.8879 | 0.597326 | 0.0589307 |
| SA 6h | 64.6014 | 13.2259 | 0.820491 | 0.0965884 |
| Me-JA 6h | 64.2506 | 4.99302 | 0.850084 | 0.0660331 |
Source Transcript PGSC0003DMT400078874 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G45290.2 | +2 | 3e-37 | 147 | 84/157 (54%) | RING/U-box superfamily protein | chr5:18350011-18352092 REVERSE LENGTH=546 |
