Probe CUST_3707_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_3707_PI426222305 | JHI_St_60k_v1 | DMT400064185 | AGGGGAAAGTGAAAGTTTGTCCGATGATTCCTTCCTCCTTTTGTTATTTTCTCCTTTTTG |
All Microarray Probes Designed to Gene DMG400024936
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_3707_PI426222305 | JHI_St_60k_v1 | DMT400064185 | AGGGGAAAGTGAAAGTTTGTCCGATGATTCCTTCCTCCTTTTGTTATTTTCTCCTTTTTG |
| CUST_3459_PI426222305 | JHI_St_60k_v1 | DMT400064186 | AGGGGAAAGTGAAAGTTTGTCCGATGATTCCTTCCTCCTTTTGTTATTTTCTCCTTTTTG |
Microarray Signals from CUST_3707_PI426222305
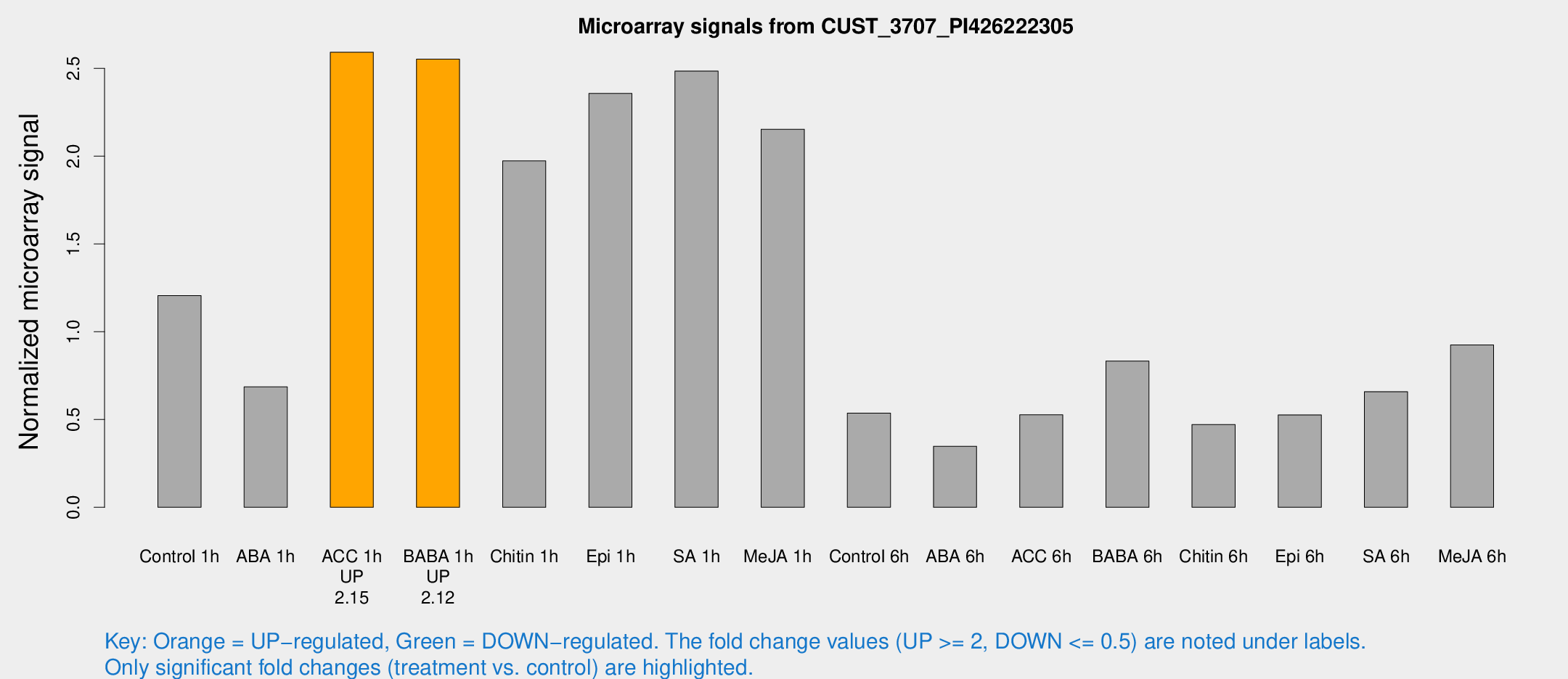
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1123.73 | 65.0055 | 1.2054 | 0.0697101 |
| ABA 1h | 663.345 | 272.825 | 0.686188 | 0.254819 |
| ACC 1h | 2490.99 | 159.638 | 2.5922 | 0.204238 |
| BABA 1h | 2413.14 | 508.356 | 2.55249 | 0.343958 |
| Chitin 1h | 1654.55 | 95.6574 | 1.97297 | 0.201717 |
| Epi 1h | 1953.03 | 298.392 | 2.35734 | 0.364 |
| SA 1h | 2497.12 | 531.891 | 2.48502 | 0.531984 |
| Me-JA 1h | 1642.02 | 95.0697 | 2.15368 | 0.255131 |
| Control 6h | 613.324 | 215.522 | 0.536037 | 0.248659 |
| ABA 6h | 370.064 | 96.248 | 0.347747 | 0.111156 |
| ACC 6h | 589.893 | 131.453 | 0.527189 | 0.0374679 |
| BABA 6h | 896.521 | 163.734 | 0.833041 | 0.118092 |
| Chitin 6h | 480.355 | 75.4422 | 0.471427 | 0.0648735 |
| Epi 6h | 559.007 | 56.0811 | 0.526369 | 0.0366382 |
| SA 6h | 610.646 | 45.452 | 0.658527 | 0.0382932 |
| Me-JA 6h | 887.498 | 151.924 | 0.925182 | 0.120424 |
Source Transcript PGSC0003DMT400064185 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G67420.1 | +3 | 1e-79 | 247 | 134/226 (59%) | LOB domain-containing protein 37 | chr5:26904576-26905415 REVERSE LENGTH=250 |
