Probe CUST_36935_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_36935_PI426222305 | JHI_St_60k_v1 | DMT400067456 | TATTATGGCAGCCCTGATTGCTATGGAACCCCTAGTGGAGCATGTGGATTTGGTGAATAT |
All Microarray Probes Designed to Gene DMG400026220
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_36901_PI426222305 | JHI_St_60k_v1 | DMT400067455 | GATGTTTTACCTGATGAATGGAAGGCTGGCATTGCTTATGACACATACCTTATGCTAGAC |
| CUST_36935_PI426222305 | JHI_St_60k_v1 | DMT400067456 | TATTATGGCAGCCCTGATTGCTATGGAACCCCTAGTGGAGCATGTGGATTTGGTGAATAT |
Microarray Signals from CUST_36935_PI426222305
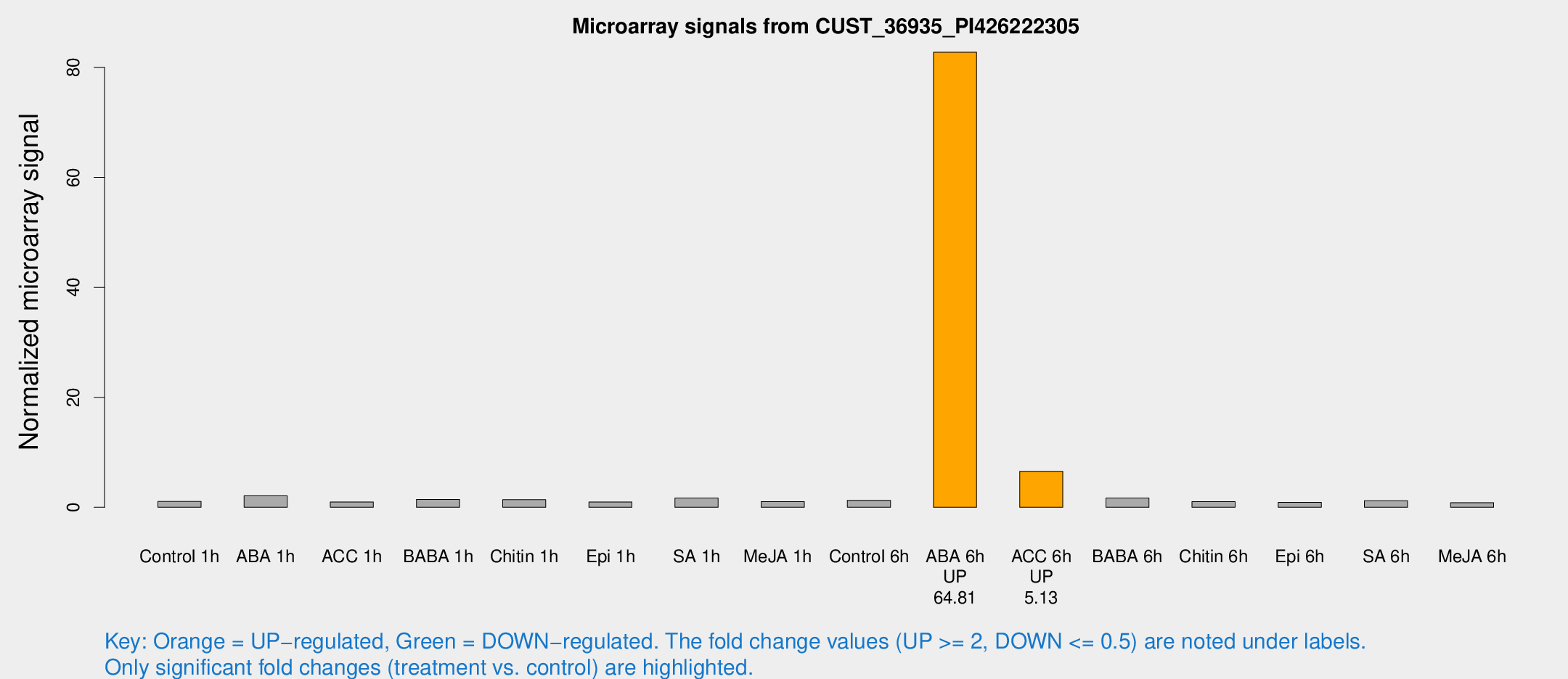
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6.98924 | 2.95114 | 1.05833 | 0.510792 |
| ABA 1h | 13.4994 | 4.52161 | 2.10324 | 1.20509 |
| ACC 1h | 6.13638 | 3.41213 | 0.953965 | 0.525761 |
| BABA 1h | 10.8861 | 5.41645 | 1.44294 | 0.674032 |
| Chitin 1h | 8.01889 | 2.95824 | 1.39407 | 0.560497 |
| Epi 1h | 4.99074 | 2.8981 | 0.951577 | 0.536683 |
| SA 1h | 12.3936 | 3.99563 | 1.69301 | 0.809075 |
| Me-JA 1h | 5.19362 | 3.0202 | 1.0336 | 0.590681 |
| Control 6h | 9.66306 | 4.44295 | 1.27708 | 0.587494 |
| ABA 6h | 585.955 | 135.377 | 82.7628 | 20.0603 |
| ACC 6h | 50.5879 | 13.5991 | 6.55348 | 1.39758 |
| BABA 6h | 18.0358 | 11.8546 | 1.67715 | 1.37576 |
| Chitin 6h | 6.90703 | 3.3664 | 1.02046 | 0.521192 |
| Epi 6h | 6.34379 | 3.5299 | 0.897392 | 0.497191 |
| SA 6h | 7.86977 | 3.23355 | 1.19106 | 0.570786 |
| Me-JA 6h | 5.21085 | 3.0201 | 0.844324 | 0.486496 |
Source Transcript PGSC0003DMT400067456 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G17030.1 | +2 | 1e-93 | 282 | 134/250 (54%) | expansin-like B1 | chr4:9581817-9583181 REVERSE LENGTH=250 |
