Probe CUST_36033_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_36033_PI426222305 | JHI_St_60k_v1 | DMT400042790 | CTGAAAGAGCTGCTAGATTTTCTTGTTTTGCTACAAATTTGGAAAGTGGACAACTTTCAA |
All Microarray Probes Designed to Gene DMG400016608
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_35971_PI426222305 | JHI_St_60k_v1 | DMT400042791 | CTGAAAGAGCTGCTAGATTTTCTTGTTTTGCTACAAATTTGGAAAGTGGACAACTTTCAA |
| CUST_36033_PI426222305 | JHI_St_60k_v1 | DMT400042790 | CTGAAAGAGCTGCTAGATTTTCTTGTTTTGCTACAAATTTGGAAAGTGGACAACTTTCAA |
Microarray Signals from CUST_36033_PI426222305
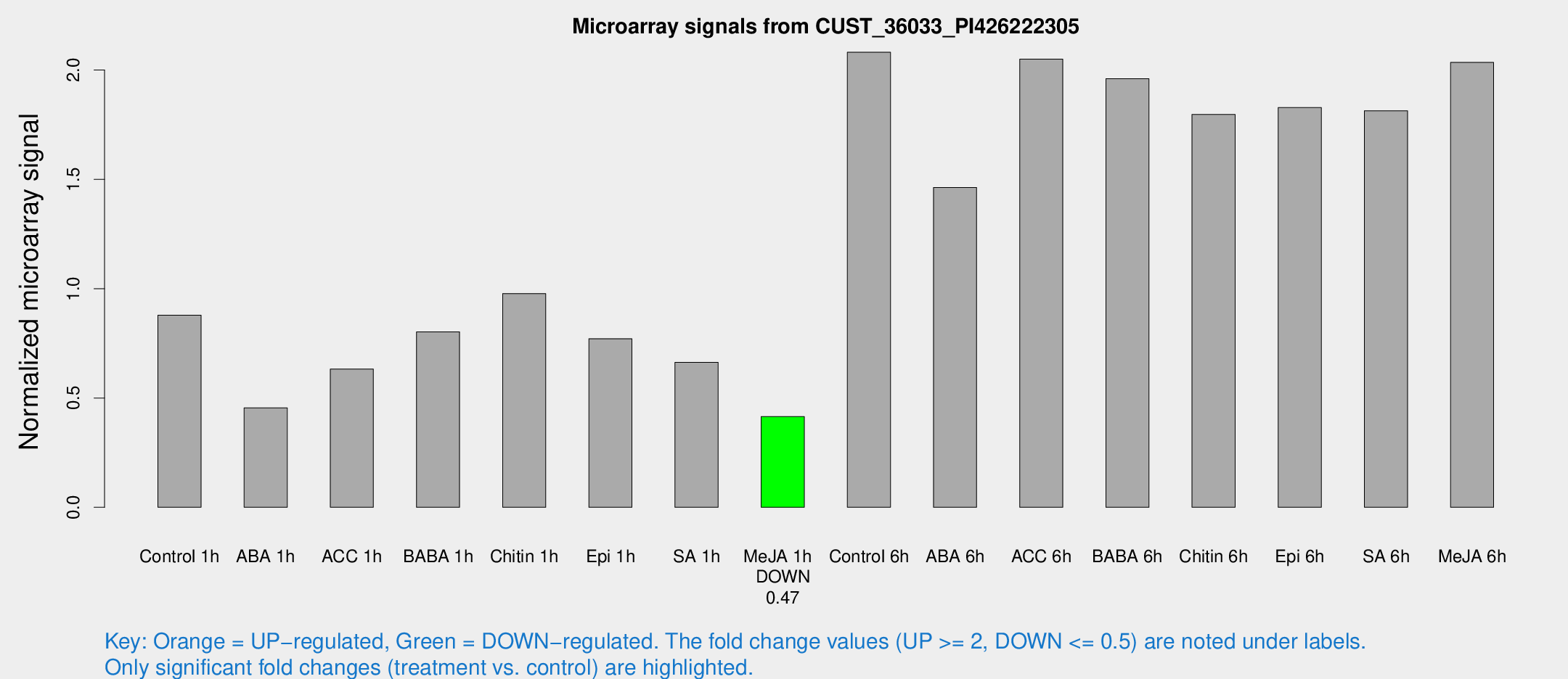
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 260.925 | 15.462 | 0.879022 | 0.0520496 |
| ABA 1h | 123.133 | 22.5261 | 0.454933 | 0.0494644 |
| ACC 1h | 202.563 | 41.5448 | 0.632477 | 0.0988641 |
| BABA 1h | 235.7 | 36.6461 | 0.802511 | 0.0541115 |
| Chitin 1h | 261.681 | 15.6951 | 0.977542 | 0.0579702 |
| Epi 1h | 203.462 | 31.5648 | 0.770964 | 0.123251 |
| SA 1h | 203.858 | 20.4734 | 0.662703 | 0.0399087 |
| Me-JA 1h | 101.499 | 10.3529 | 0.41507 | 0.0281772 |
| Control 6h | 720.096 | 239.47 | 2.08149 | 0.750826 |
| ABA 6h | 478.182 | 84.1485 | 1.4631 | 0.218676 |
| ACC 6h | 759.128 | 192.61 | 2.04959 | 0.517865 |
| BABA 6h | 691.68 | 152.39 | 1.96047 | 0.449397 |
| Chitin 6h | 584.943 | 95.5651 | 1.79689 | 0.34352 |
| Epi 6h | 643.509 | 131.399 | 1.82848 | 0.594116 |
| SA 6h | 622.623 | 210.29 | 1.81389 | 1.16805 |
| Me-JA 6h | 659.366 | 173.623 | 2.03483 | 0.524773 |
Source Transcript PGSC0003DMT400042790 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G07340.1 | +1 | 4e-69 | 235 | 161/353 (46%) | basic helix-loop-helix (bHLH) DNA-binding superfamily protein | chr3:2341188-2343288 REVERSE LENGTH=456 |
