Probe CUST_36030_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_36030_PI426222305 | JHI_St_60k_v1 | DMT400056316 | GGTTTCCTCAAGGTTTTCCTCCAGAATGTAGAAGATTTTGAGTGGGCCAAGTAGTTTTTT |
All Microarray Probes Designed to Gene DMG400021877
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_36030_PI426222305 | JHI_St_60k_v1 | DMT400056316 | GGTTTCCTCAAGGTTTTCCTCCAGAATGTAGAAGATTTTGAGTGGGCCAAGTAGTTTTTT |
Microarray Signals from CUST_36030_PI426222305
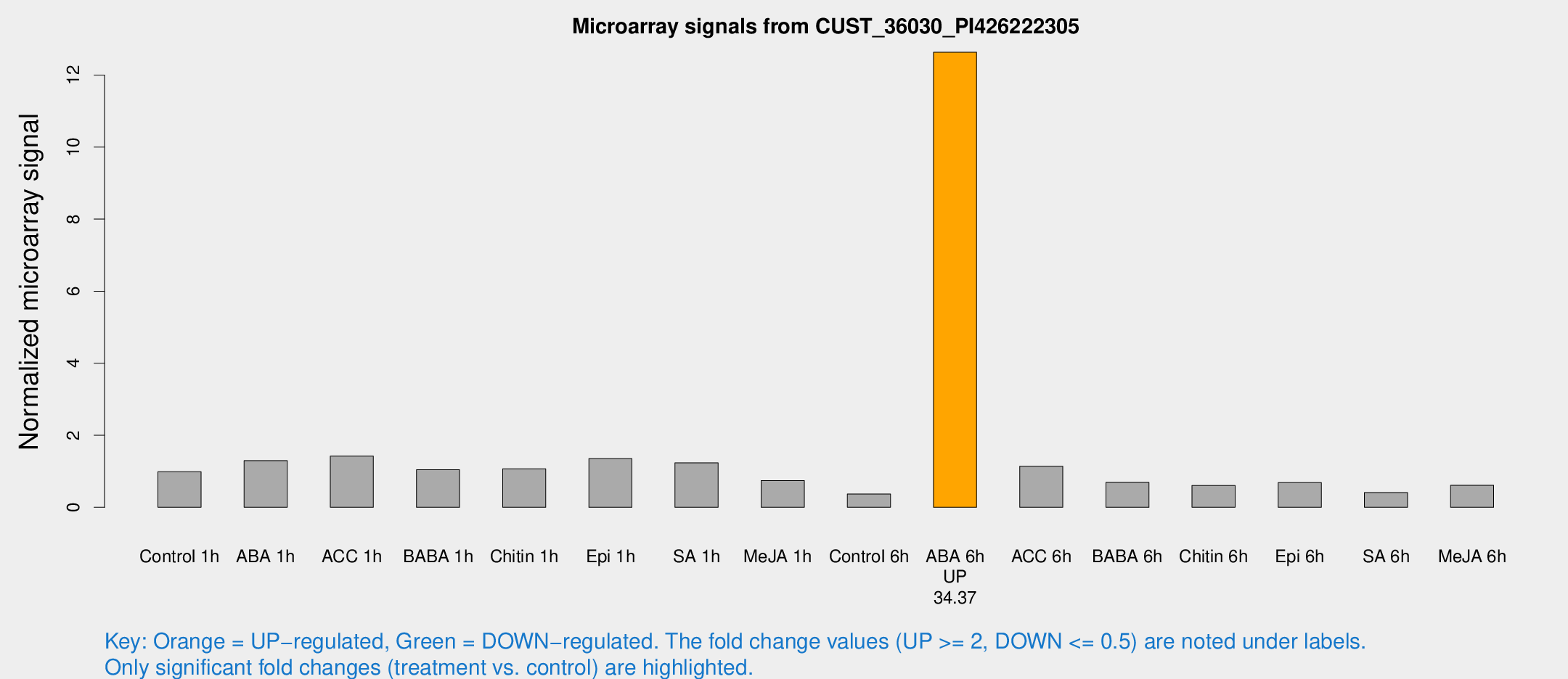
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 2254.27 | 465.359 | 0.98719 | 0.156708 |
| ABA 1h | 2514.88 | 185.03 | 1.29754 | 0.188067 |
| ACC 1h | 3417.38 | 861.884 | 1.42298 | 0.288018 |
| BABA 1h | 2258.25 | 407.088 | 1.04202 | 0.121311 |
| Chitin 1h | 2121.87 | 250.917 | 1.06734 | 0.194895 |
| Epi 1h | 2557.16 | 148.065 | 1.35201 | 0.0780802 |
| SA 1h | 2808.36 | 353.641 | 1.23512 | 0.125448 |
| Me-JA 1h | 1321.12 | 76.574 | 0.74048 | 0.0915193 |
| Control 6h | 918.907 | 281.505 | 0.367543 | 0.120895 |
| ABA 6h | 29657.1 | 3477.52 | 12.6343 | 0.780144 |
| ACC 6h | 2876.92 | 265.555 | 1.14085 | 0.10325 |
| BABA 6h | 1967.58 | 805.936 | 0.691109 | 0.271922 |
| Chitin 6h | 1431.3 | 202.676 | 0.603604 | 0.10876 |
| Epi 6h | 1706.87 | 193.429 | 0.684969 | 0.0469533 |
| SA 6h | 899.294 | 144.558 | 0.407057 | 0.0458771 |
| Me-JA 6h | 1408.03 | 357.347 | 0.61187 | 0.159503 |
Source Transcript PGSC0003DMT400056316 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G14130.1 | +3 | 1e-151 | 432 | 201/276 (73%) | xyloglucan endotransglucosylase/hydrolase 15 | chr4:8137161-8138196 REVERSE LENGTH=289 |
