Probe CUST_36023_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_36023_PI426222305 | JHI_St_60k_v1 | DMT400053987 | GTGGATCTTCAGAAGTGAATCTAAAAGCTTTTGTTCAAGAAGTTGGCAAAAGTTGTTGAA |
All Microarray Probes Designed to Gene DMG400020944
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_36023_PI426222305 | JHI_St_60k_v1 | DMT400053987 | GTGGATCTTCAGAAGTGAATCTAAAAGCTTTTGTTCAAGAAGTTGGCAAAAGTTGTTGAA |
Microarray Signals from CUST_36023_PI426222305
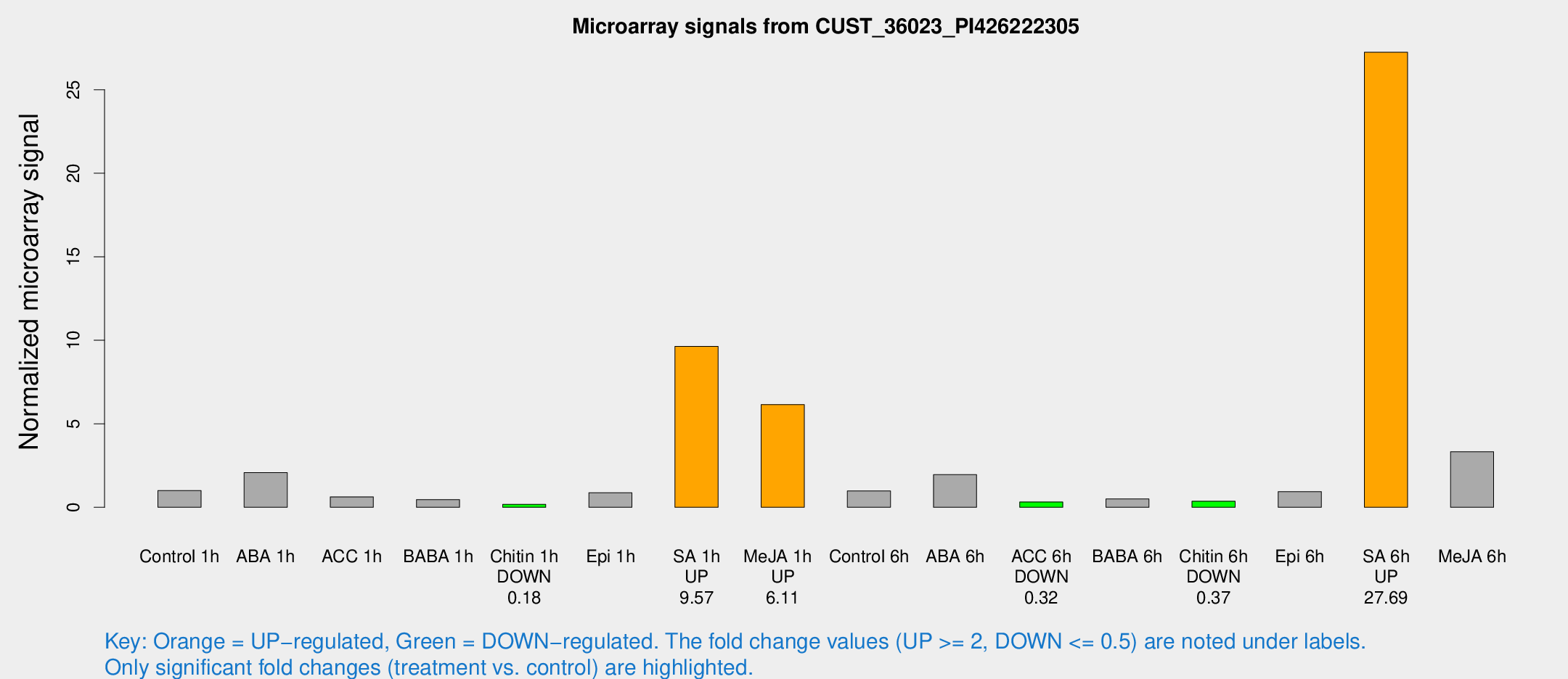
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 90.2695 | 15.6661 | 1.00666 | 0.104063 |
| ABA 1h | 173.09 | 43.6191 | 2.08097 | 0.538572 |
| ACC 1h | 61.8329 | 19.9251 | 0.626904 | 0.173844 |
| BABA 1h | 52.419 | 21.0781 | 0.460297 | 0.305637 |
| Chitin 1h | 16.1237 | 5.56339 | 0.178152 | 0.0750359 |
| Epi 1h | 78.1478 | 33.4193 | 0.872783 | 0.418353 |
| SA 1h | 961.621 | 323.348 | 9.63702 | 2.6857 |
| Me-JA 1h | 502.8 | 191.62 | 6.14963 | 1.84309 |
| Control 6h | 88.0872 | 14.278 | 0.983939 | 0.170385 |
| ABA 6h | 187.192 | 35.4292 | 1.95389 | 0.249929 |
| ACC 6h | 32.7842 | 5.10669 | 0.319009 | 0.104847 |
| BABA 6h | 55.2114 | 17.4075 | 0.501599 | 0.229074 |
| Chitin 6h | 34.0574 | 4.7063 | 0.3651 | 0.0510212 |
| Epi 6h | 91.826 | 6.92402 | 0.934317 | 0.0755241 |
| SA 6h | 2532.31 | 654.74 | 27.2435 | 4.97537 |
| Me-JA 6h | 314.371 | 92.1831 | 3.33086 | 1.29452 |
Source Transcript PGSC0003DMT400053987 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G14090.1 | +2 | 8e-42 | 146 | 66/107 (62%) | UDP-Glycosyltransferase superfamily protein | chr4:8122434-8123804 REVERSE LENGTH=456 |
