Probe CUST_35538_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_35538_PI426222305 | JHI_St_60k_v1 | DMT400032519 | CAGTTGCAGTTGTGCTACAGCCTACAATTGGTTTCTAGTTTTATTTTCTTCTCATTTTAG |
All Microarray Probes Designed to Gene DMG400012491
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_35538_PI426222305 | JHI_St_60k_v1 | DMT400032519 | CAGTTGCAGTTGTGCTACAGCCTACAATTGGTTTCTAGTTTTATTTTCTTCTCATTTTAG |
| CUST_35526_PI426222305 | JHI_St_60k_v1 | DMT400032518 | TTGACAACAAAACAAACACAAACATTGCTGAAAAGCACACTACCCGACTTTGGTCCCCTG |
Microarray Signals from CUST_35538_PI426222305
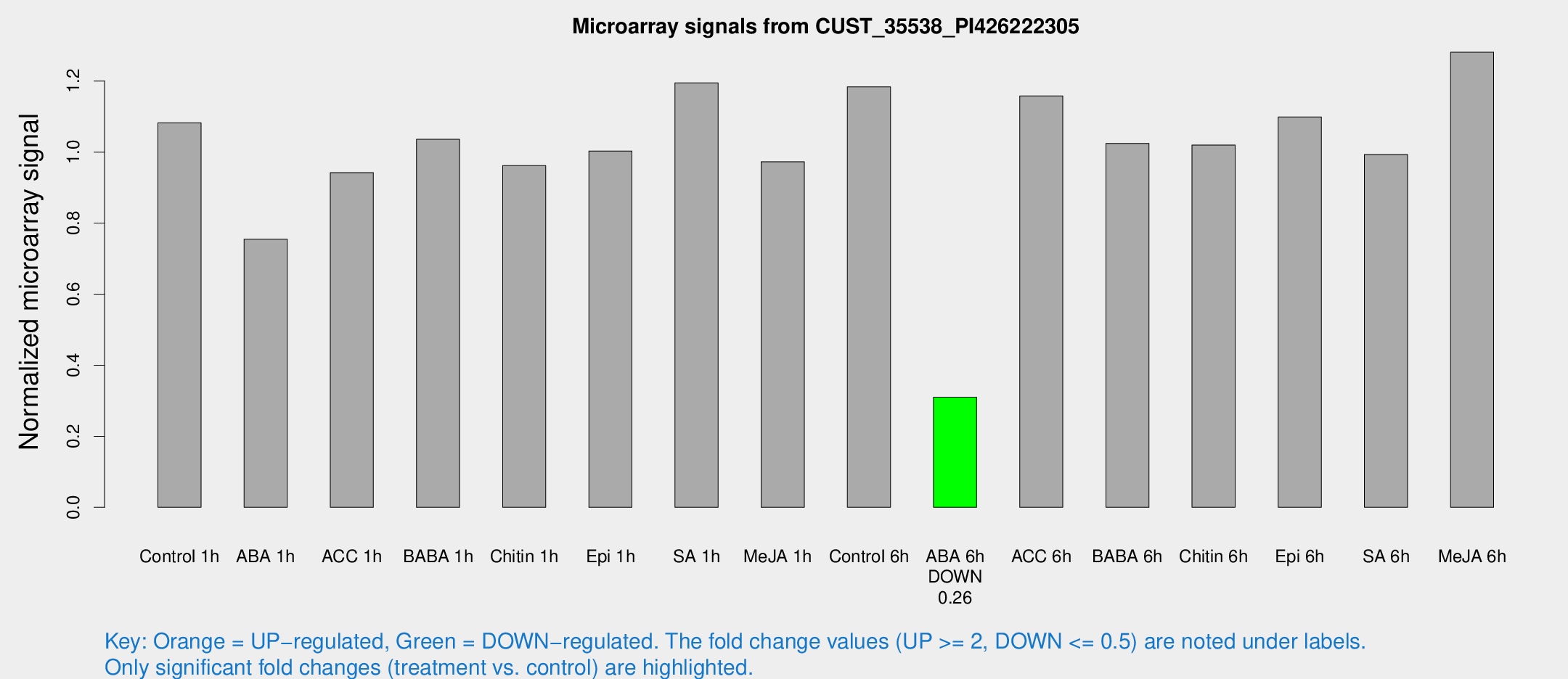
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 761.131 | 144.424 | 1.08252 | 0.147738 |
| ABA 1h | 455.868 | 27.3705 | 0.755054 | 0.0727605 |
| ACC 1h | 695.826 | 151.189 | 0.942092 | 0.16656 |
| BABA 1h | 705.292 | 135.695 | 1.03593 | 0.11868 |
| Chitin 1h | 593.19 | 53.3489 | 0.962009 | 0.0557852 |
| Epi 1h | 590.53 | 34.2487 | 1.00314 | 0.0581621 |
| SA 1h | 855.051 | 125.72 | 1.19479 | 0.140591 |
| Me-JA 1h | 552.307 | 79.7067 | 0.972916 | 0.113106 |
| Control 6h | 905.985 | 270.116 | 1.18395 | 0.332465 |
| ABA 6h | 224.485 | 13.4202 | 0.310262 | 0.0185336 |
| ACC 6h | 925.52 | 148.098 | 1.15804 | 0.0670406 |
| BABA 6h | 785.174 | 64.1314 | 1.02464 | 0.0634026 |
| Chitin 6h | 740.561 | 43.0399 | 1.02005 | 0.0591166 |
| Epi 6h | 875.041 | 164.262 | 1.09878 | 0.285252 |
| SA 6h | 681.227 | 97.8883 | 0.993145 | 0.10363 |
| Me-JA 6h | 897.213 | 155.068 | 1.28122 | 0.1359 |
Source Transcript PGSC0003DMT400032519 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G67360.1 | +2 | 2e-154 | 415 | 266/568 (47%) | Subtilase family protein | chr5:26872192-26874465 REVERSE LENGTH=757 |
