Probe CUST_35448_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_35448_PI426222305 | JHI_St_60k_v1 | DMT400005823 | CTGTCAAGCCAGTTAGCATCATAATCTGTATTGTCTTGTGTATATGTGTAATCTATAGGT |
All Microarray Probes Designed to Gene DMG401002270
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_35448_PI426222305 | JHI_St_60k_v1 | DMT400005823 | CTGTCAAGCCAGTTAGCATCATAATCTGTATTGTCTTGTGTATATGTGTAATCTATAGGT |
| CUST_35536_PI426222305 | JHI_St_60k_v1 | DMT400005803 | AAAGTGAAGATTGATACTGAATTTGTGGAGGTTATGGCTTTCAGGCATGTTCATGATGTG |
Microarray Signals from CUST_35448_PI426222305
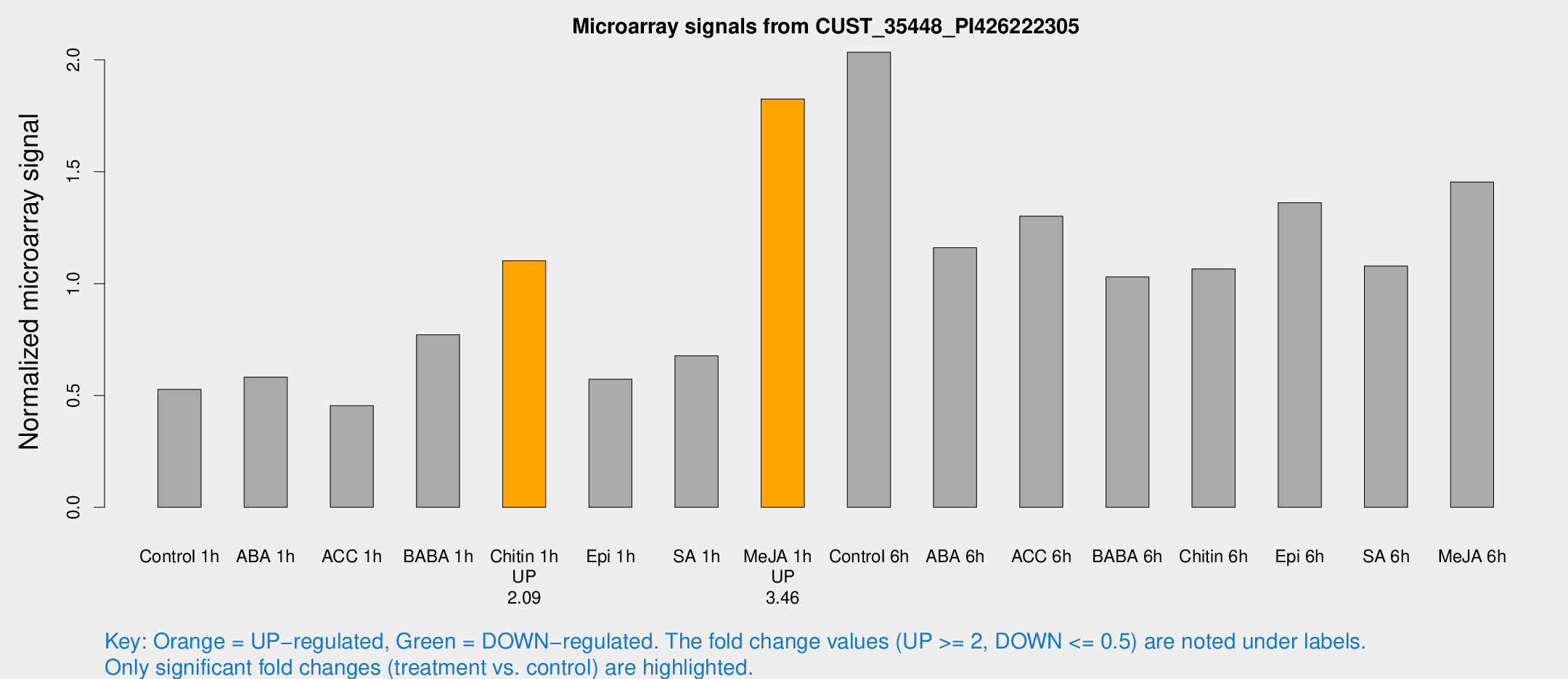
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 4774.83 | 365.71 | 0.527233 | 0.0419825 |
| ABA 1h | 4810.71 | 949.907 | 0.581887 | 0.0902382 |
| ACC 1h | 4296.7 | 611.972 | 0.454612 | 0.0915101 |
| BABA 1h | 7105.31 | 1585.46 | 0.77092 | 0.11964 |
| Chitin 1h | 8937.1 | 516.798 | 1.10227 | 0.063641 |
| Epi 1h | 4581.52 | 727.942 | 0.572423 | 0.0837351 |
| SA 1h | 6351.17 | 760.102 | 0.677318 | 0.0758591 |
| Me-JA 1h | 13683 | 1846.96 | 1.82515 | 0.119468 |
| Control 6h | 19350.5 | 4394.99 | 2.03416 | 0.301443 |
| ABA 6h | 11423.4 | 1964.07 | 1.16055 | 0.129781 |
| ACC 6h | 13723.8 | 1933.4 | 1.30168 | 0.0879869 |
| BABA 6h | 10449.4 | 894.363 | 1.02945 | 0.0713644 |
| Chitin 6h | 10231.8 | 591.842 | 1.06572 | 0.0800809 |
| Epi 6h | 14265.6 | 2508.92 | 1.36108 | 0.319389 |
| SA 6h | 10201.9 | 2300.81 | 1.07882 | 0.156203 |
| Me-JA 6h | 13464.3 | 2356.63 | 1.45367 | 0.143333 |
Source Transcript PGSC0003DMT400005823 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G47240.1 | +1 | 3e-25 | 105 | 65/140 (46%) | nudix hydrolase homolog 8 | chr5:19183806-19185467 FORWARD LENGTH=369 |
