Probe CUST_35257_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_35257_PI426222305 | JHI_St_60k_v1 | DMT400006089 | GCAAACAGATTGCCAGTCTCAGGTGAGTTCCTCAACCTTCTCTTCTTTTCTCCTTTTTTC |
All Microarray Probes Designed to Gene DMG400002366
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_35257_PI426222305 | JHI_St_60k_v1 | DMT400006089 | GCAAACAGATTGCCAGTCTCAGGTGAGTTCCTCAACCTTCTCTTCTTTTCTCCTTTTTTC |
| CUST_35269_PI426222305 | JHI_St_60k_v1 | DMT400006092 | CGGGCCTTTGCCCTTCTAGTTCTACTATATTATAAAAGAGAGTTTAGGCACAAATATAAT |
Microarray Signals from CUST_35257_PI426222305
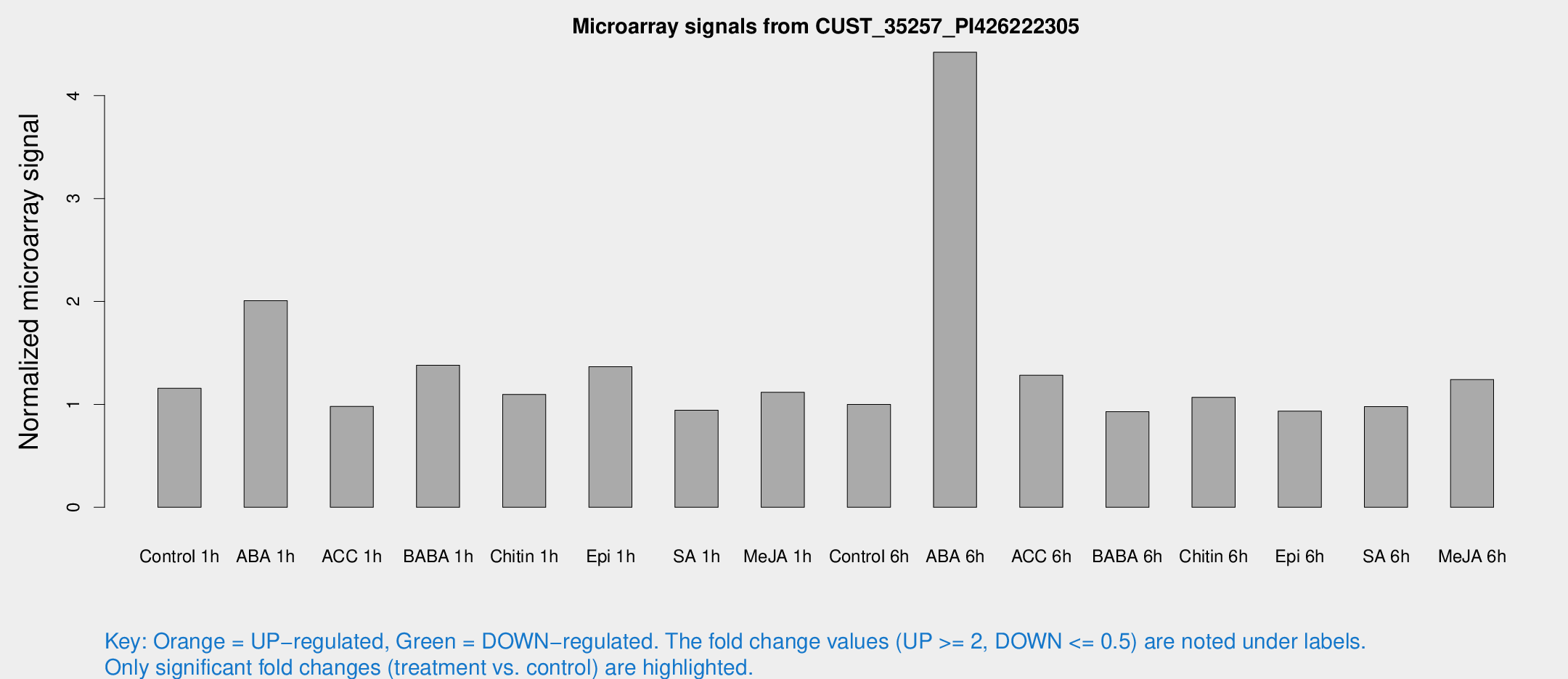
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 7.16291 | 3.01887 | 1.15697 | 0.553832 |
| ABA 1h | 11.2521 | 2.97287 | 2.00817 | 0.670159 |
| ACC 1h | 5.87542 | 3.4218 | 0.980731 | 0.567916 |
| BABA 1h | 8.1762 | 3.18403 | 1.38062 | 0.612084 |
| Chitin 1h | 5.74369 | 3.00789 | 1.0969 | 0.579126 |
| Epi 1h | 7.43352 | 2.99174 | 1.36631 | 0.639609 |
| SA 1h | 5.66881 | 3.01749 | 0.942917 | 0.508077 |
| Me-JA 1h | 5.29948 | 3.08526 | 1.11753 | 0.647027 |
| Control 6h | 5.8636 | 3.10312 | 1.00034 | 0.537675 |
| ABA 6h | 28.4058 | 5.73368 | 4.422 | 0.846401 |
| ACC 6h | 9.18539 | 3.82961 | 1.28306 | 0.608757 |
| BABA 6h | 6.04392 | 3.46058 | 0.929406 | 0.530834 |
| Chitin 6h | 6.69402 | 3.43025 | 1.06908 | 0.562243 |
| Epi 6h | 6.15813 | 3.59577 | 0.934372 | 0.541458 |
| SA 6h | 5.61456 | 3.25588 | 0.977885 | 0.566783 |
| Me-JA 6h | 7.77656 | 3.12874 | 1.24175 | 0.581356 |
Source Transcript PGSC0003DMT400006089 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G62020.1 | +1 | 1e-21 | 92 | 41/50 (82%) | Coatomer, alpha subunit | chr1:22919814-22923728 FORWARD LENGTH=1216 |
