Probe CUST_3484_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_3484_PI426222305 | JHI_St_60k_v1 | DMT400010274 | GTTGGTAGCGTGGAAGACTTAATAGAATTTCCAAAAGACTGACATTGAGTGAGAATAATA |
All Microarray Probes Designed to Gene DMG400004017
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_3484_PI426222305 | JHI_St_60k_v1 | DMT400010274 | GTTGGTAGCGTGGAAGACTTAATAGAATTTCCAAAAGACTGACATTGAGTGAGAATAATA |
| CUST_3706_PI426222305 | JHI_St_60k_v1 | DMT400010275 | GTTGGTAGCGTGGAAGACTTAATAGAATTTCCAAAAGACTGACATTGAGTGAGAATAATA |
Microarray Signals from CUST_3484_PI426222305
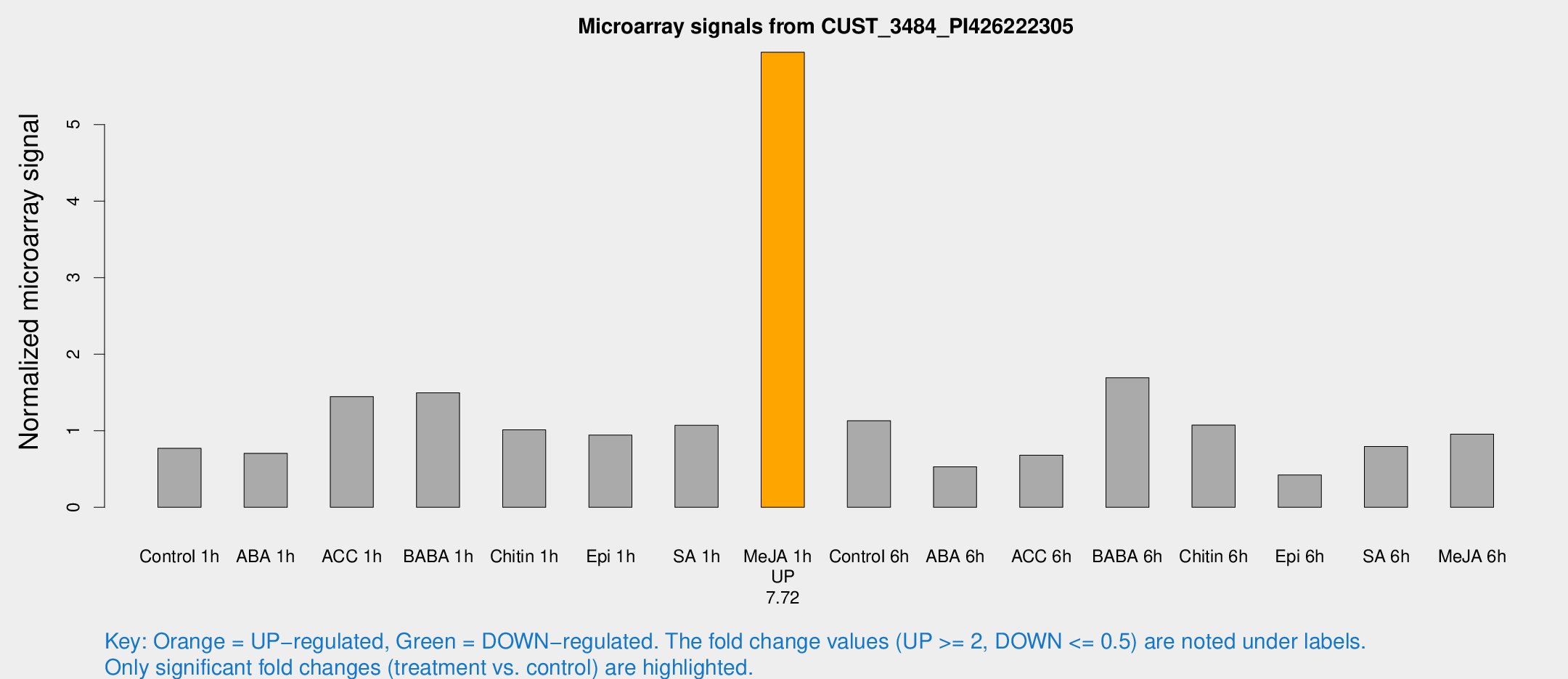
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 105.399 | 11.3632 | 0.770569 | 0.0516615 |
| ABA 1h | 94.5198 | 33.1982 | 0.705607 | 0.2082 |
| ACC 1h | 204.389 | 26.9614 | 1.44716 | 0.0949489 |
| BABA 1h | 199.85 | 30.1335 | 1.49658 | 0.388158 |
| Chitin 1h | 137.216 | 44.5402 | 1.01222 | 0.331126 |
| Epi 1h | 112.022 | 13.2156 | 0.943517 | 0.109206 |
| SA 1h | 174.886 | 69.0213 | 1.07075 | 0.544692 |
| Me-JA 1h | 656.383 | 38.103 | 5.94593 | 0.660343 |
| Control 6h | 205.272 | 82.6424 | 1.13034 | 0.737998 |
| ABA 6h | 99.7028 | 52.8999 | 0.529962 | 0.367923 |
| ACC 6h | 126.573 | 56.3648 | 0.681591 | 0.198964 |
| BABA 6h | 269.097 | 62.7407 | 1.6934 | 0.330688 |
| Chitin 6h | 162.367 | 37.2763 | 1.07448 | 0.237518 |
| Epi 6h | 69.9396 | 17.5384 | 0.422995 | 0.171423 |
| SA 6h | 118.471 | 38.0639 | 0.794593 | 0.316793 |
| Me-JA 6h | 131.23 | 18.2035 | 0.955709 | 0.0620096 |
Source Transcript PGSC0003DMT400010274 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G42930.1 | +3 | 1e-130 | 395 | 223/466 (48%) | alpha/beta-Hydrolases superfamily protein | chr5:17210738-17214152 REVERSE LENGTH=467 |
