Probe CUST_34330_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_34330_PI426222305 | JHI_St_60k_v1 | DMT400082751 | CTACCTAAACATTTTAAGCACAACAACTTCTCTAGCTTTATCCGCCAGCTTAATACCTAT |
All Microarray Probes Designed to Gene DMG400032793
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_34326_PI426222305 | JHI_St_60k_v1 | DMT400082752 | CTACCTAAACATTTTAAGCACAACAACTTCTCTAGCTTTATCCGCCAGCTTAATACCTAT |
| CUST_34330_PI426222305 | JHI_St_60k_v1 | DMT400082751 | CTACCTAAACATTTTAAGCACAACAACTTCTCTAGCTTTATCCGCCAGCTTAATACCTAT |
Microarray Signals from CUST_34330_PI426222305
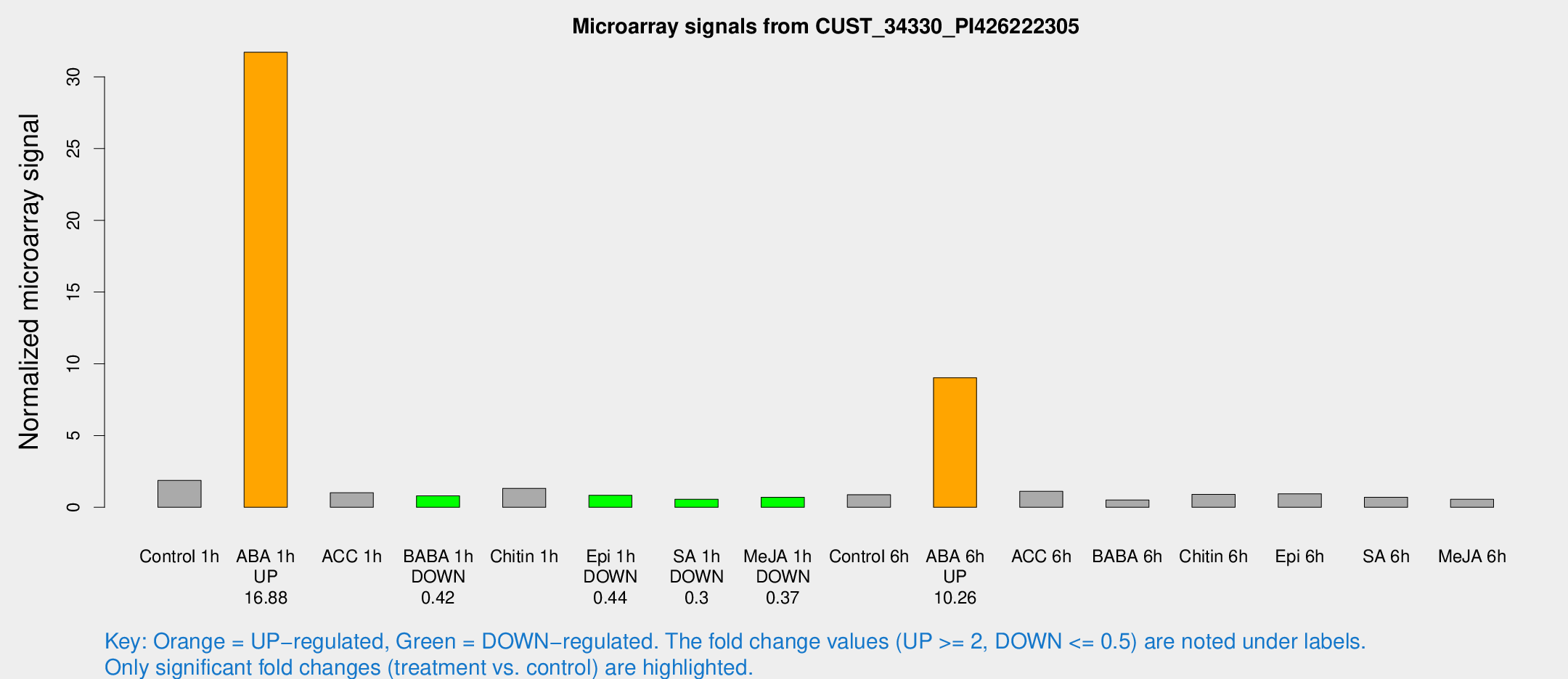
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 21.6678 | 4.10651 | 1.87866 | 0.309592 |
| ABA 1h | 323.701 | 65.2872 | 31.714 | 5.96483 |
| ACC 1h | 12.5464 | 3.38889 | 1.01378 | 0.356166 |
| BABA 1h | 9.10988 | 3.11542 | 0.79793 | 0.319301 |
| Chitin 1h | 13.4257 | 3.02542 | 1.32206 | 0.311424 |
| Epi 1h | 8.29539 | 2.92398 | 0.834149 | 0.328756 |
| SA 1h | 6.72235 | 2.95048 | 0.560831 | 0.273059 |
| Me-JA 1h | 6.56102 | 3.01931 | 0.698161 | 0.342896 |
| Control 6h | 10.591 | 3.07309 | 0.880408 | 0.319337 |
| ABA 6h | 112.344 | 24.164 | 9.03172 | 1.91397 |
| ACC 6h | 15.3604 | 4.01422 | 1.11095 | 0.323352 |
| BABA 6h | 6.25182 | 3.39541 | 0.503376 | 0.274706 |
| Chitin 6h | 12.6994 | 5.38103 | 0.904234 | 0.459983 |
| Epi 6h | 13.1241 | 3.75694 | 0.941669 | 0.406826 |
| SA 6h | 8.36667 | 3.22469 | 0.701968 | 0.321505 |
| Me-JA 6h | 6.24348 | 3.03569 | 0.554274 | 0.281055 |
Source Transcript PGSC0003DMT400082751 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G26150.1 | +3 | 2e-57 | 125 | 53/75 (71%) | heat shock transcription factor A2 | chr2:11135856-11137217 FORWARD LENGTH=345 |
