Probe CUST_34162_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_34162_PI426222305 | JHI_St_60k_v1 | DMT400033096 | GAGTCCAGAGAAAGAAGATGCTTCTCAACTGGAGTTATATTCTTTCATTGCTCATATTAC |
All Microarray Probes Designed to Gene DMG400012707
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_34162_PI426222305 | JHI_St_60k_v1 | DMT400033096 | GAGTCCAGAGAAAGAAGATGCTTCTCAACTGGAGTTATATTCTTTCATTGCTCATATTAC |
Microarray Signals from CUST_34162_PI426222305
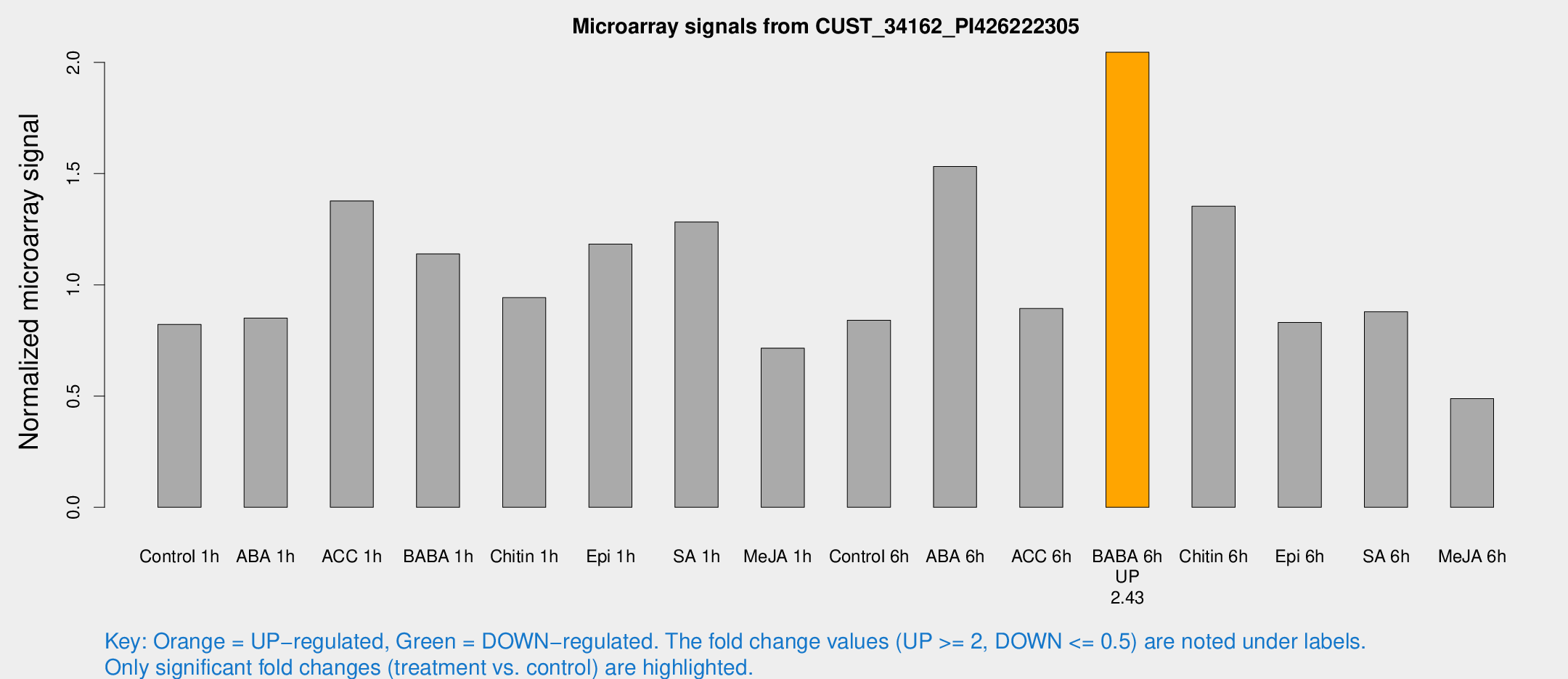
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 485.933 | 71.8531 | 0.822072 | 0.0650841 |
| ABA 1h | 438.647 | 38.0858 | 0.850655 | 0.0857512 |
| ACC 1h | 826.769 | 80.1072 | 1.37721 | 0.0797387 |
| BABA 1h | 655.155 | 111.327 | 1.13906 | 0.100977 |
| Chitin 1h | 508.307 | 88.8161 | 0.942442 | 0.13217 |
| Epi 1h | 593.8 | 34.4969 | 1.18322 | 0.0689097 |
| SA 1h | 826.239 | 212.735 | 1.28249 | 0.316548 |
| Me-JA 1h | 343.713 | 42.8289 | 0.715458 | 0.0680312 |
| Control 6h | 509.113 | 96.3592 | 0.840876 | 0.104254 |
| ABA 6h | 979.959 | 194.617 | 1.53177 | 0.219606 |
| ACC 6h | 610.665 | 97.2525 | 0.893377 | 0.117155 |
| BABA 6h | 1328.06 | 76.934 | 2.04562 | 0.11824 |
| Chitin 6h | 850.716 | 124.924 | 1.35335 | 0.150185 |
| Epi 6h | 543.409 | 31.6722 | 0.830966 | 0.0682847 |
| SA 6h | 504.359 | 29.3954 | 0.878793 | 0.126233 |
| Me-JA 6h | 319.15 | 107.593 | 0.488736 | 0.143429 |
Source Transcript PGSC0003DMT400033096 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G18570.1 | +1 | 6e-35 | 128 | 76/168 (45%) | UDP-Glycosyltransferase superfamily protein | chr2:8063429-8064841 FORWARD LENGTH=470 |
