Probe CUST_34141_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_34141_PI426222305 | JHI_St_60k_v1 | DMT400033140 | ACATGACCCAAAGCAAAACGAGATCTGCAAATGGGACGGGAATCTTTCCCTAATCGTGAT |
All Microarray Probes Designed to Gene DMG404012723
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_34141_PI426222305 | JHI_St_60k_v1 | DMT400033140 | ACATGACCCAAAGCAAAACGAGATCTGCAAATGGGACGGGAATCTTTCCCTAATCGTGAT |
| CUST_34133_PI426222305 | JHI_St_60k_v1 | DMT400033138 | TTGAGTTTCTCTCTTCAATTGTAGGGACGGGAATCTTTCCCTAATCGTGATGGCGTTGAG |
Microarray Signals from CUST_34141_PI426222305
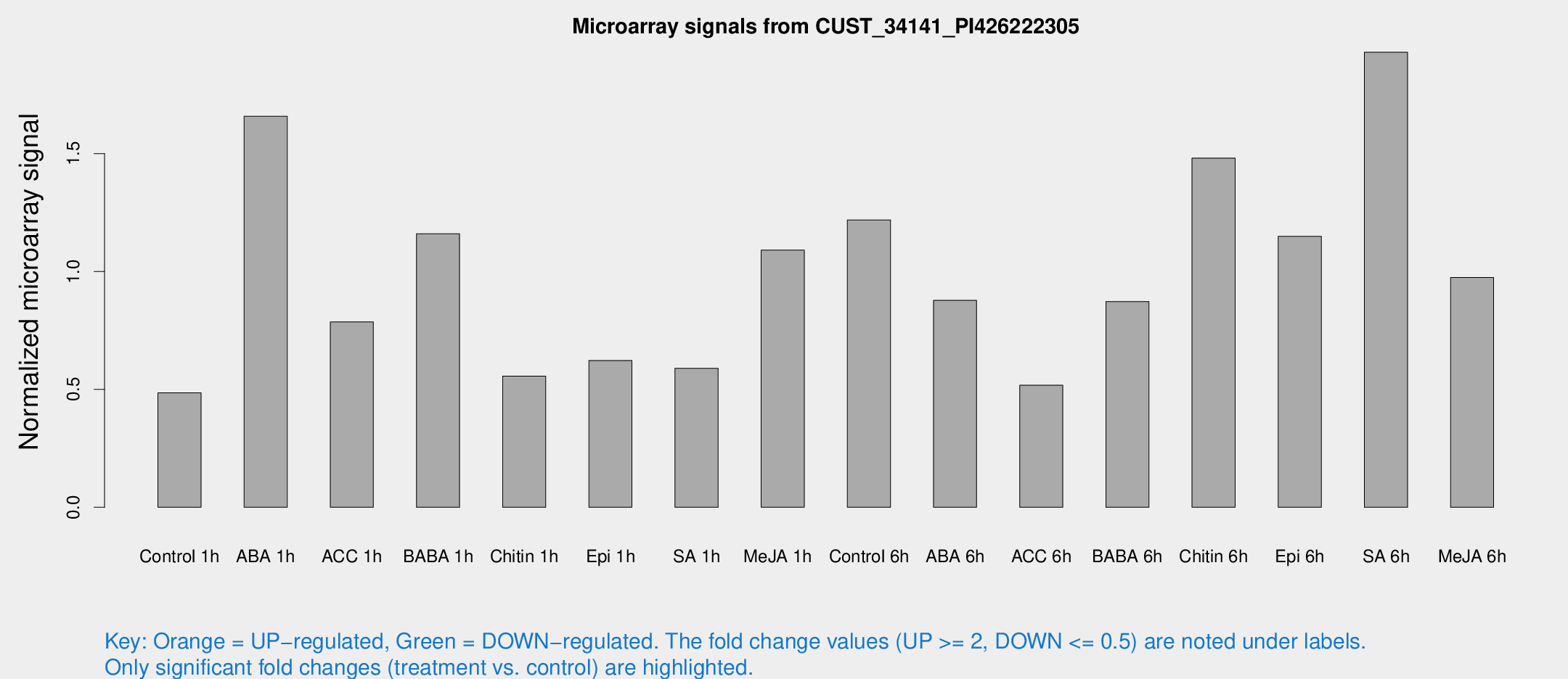
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 8.27623 | 3.57705 | 0.485469 | 0.219905 |
| ABA 1h | 26.0153 | 5.74696 | 1.65815 | 0.476227 |
| ACC 1h | 14.4614 | 4.07092 | 0.786188 | 0.253399 |
| BABA 1h | 18.7161 | 3.95333 | 1.16005 | 0.246323 |
| Chitin 1h | 9.28959 | 3.68793 | 0.55606 | 0.265795 |
| Epi 1h | 9.94866 | 3.63001 | 0.622601 | 0.271371 |
| SA 1h | 11.2492 | 3.71625 | 0.589357 | 0.255312 |
| Me-JA 1h | 14.8771 | 3.75646 | 1.09064 | 0.275874 |
| Control 6h | 28.1828 | 16.2843 | 1.21796 | 0.765182 |
| ABA 6h | 16.7178 | 4.5075 | 0.877452 | 0.244267 |
| ACC 6h | 10.6172 | 4.65472 | 0.517656 | 0.248962 |
| BABA 6h | 19.59 | 6.90463 | 0.8723 | 0.438634 |
| Chitin 6h | 26.7406 | 4.35112 | 1.48077 | 0.248445 |
| Epi 6h | 22.7174 | 4.73757 | 1.14893 | 0.250508 |
| SA 6h | 35.971 | 13.0661 | 1.92962 | 0.558975 |
| Me-JA 6h | 19.9039 | 7.249 | 0.974799 | 0.486393 |
Source Transcript PGSC0003DMT400033140 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G39620.1 | +1 | 0.0 | 809 | 409/844 (48%) | Pentatricopeptide repeat (PPR) superfamily protein | chr2:16518968-16521478 REVERSE LENGTH=836 |
