Probe CUST_33885_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_33885_PI426222305 | JHI_St_60k_v1 | DMT400012508 | CCCTATGTAGTTTGTTACAGCTATGATTGTCTACTGTGAAGGTTAATAGATGATCCAATT |
All Microarray Probes Designed to Gene DMG401004883
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_33885_PI426222305 | JHI_St_60k_v1 | DMT400012508 | CCCTATGTAGTTTGTTACAGCTATGATTGTCTACTGTGAAGGTTAATAGATGATCCAATT |
| CUST_33908_PI426222305 | JHI_St_60k_v1 | DMT400012509 | ATTGAAAAGGGTCCTCTTCATTTTCTTTCAGCTCTATTCTCCGGTGATTCCTATCTCCAT |
Microarray Signals from CUST_33885_PI426222305
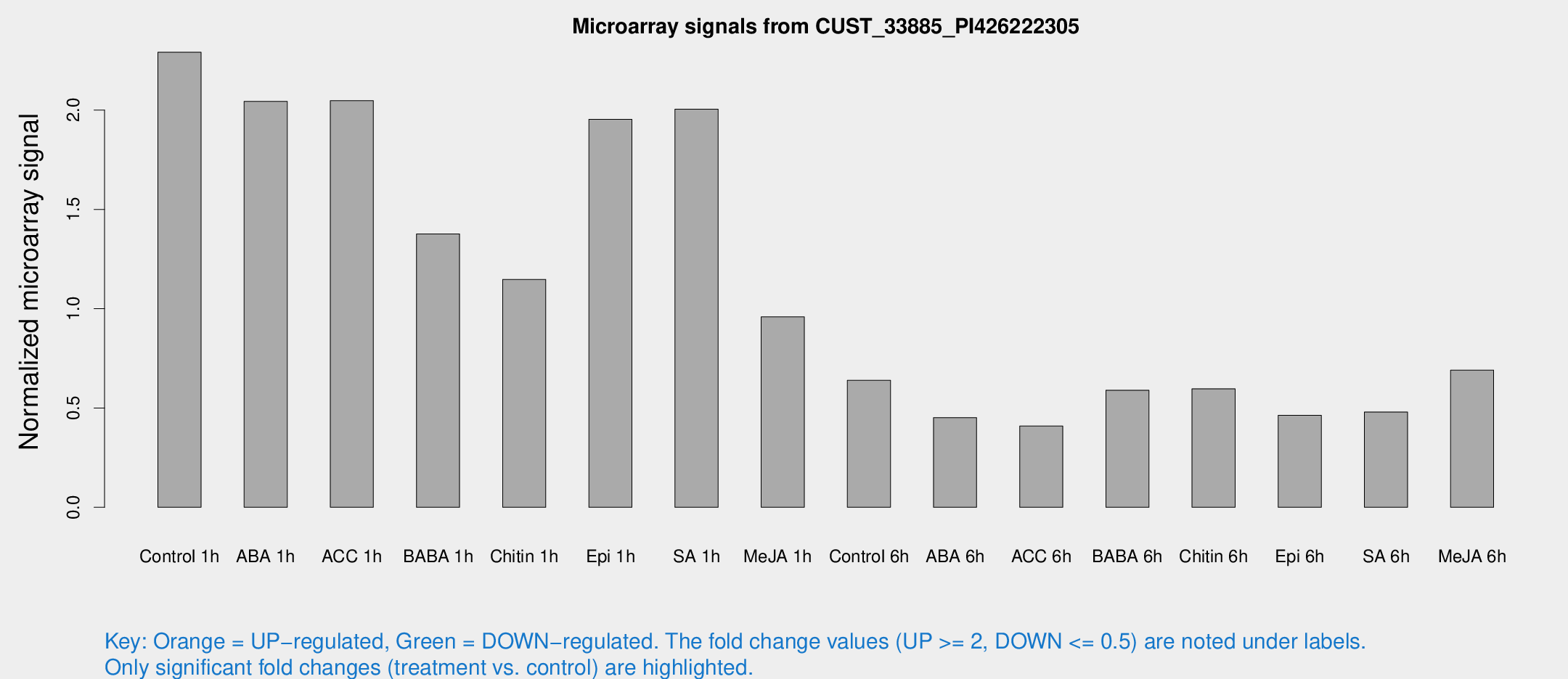
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 155.397 | 25.259 | 2.29146 | 0.227218 |
| ABA 1h | 120.597 | 11.8304 | 2.04388 | 0.351173 |
| ACC 1h | 148.277 | 34.7218 | 2.04757 | 0.397375 |
| BABA 1h | 89.0294 | 10.8675 | 1.37654 | 0.0951177 |
| Chitin 1h | 69.2646 | 8.5621 | 1.14758 | 0.0840835 |
| Epi 1h | 112.793 | 11.7646 | 1.95373 | 0.230537 |
| SA 1h | 138.854 | 20.0117 | 2.00471 | 0.381727 |
| Me-JA 1h | 52.0413 | 4.37726 | 0.959067 | 0.0951229 |
| Control 6h | 43.0263 | 5.36328 | 0.639516 | 0.0615218 |
| ABA 6h | 31.7658 | 3.73327 | 0.451744 | 0.0534049 |
| ACC 6h | 31.304 | 4.29827 | 0.409299 | 0.0867793 |
| BABA 6h | 53.0148 | 22.7565 | 0.589509 | 0.322524 |
| Chitin 6h | 45.6166 | 13.8405 | 0.596654 | 0.185276 |
| Epi 6h | 36.7569 | 9.05309 | 0.463075 | 0.168259 |
| SA 6h | 35.6487 | 10.8278 | 0.480017 | 0.135861 |
| Me-JA 6h | 46.8881 | 8.78166 | 0.690567 | 0.0889977 |
Source Transcript PGSC0003DMT400012508 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G34110.1 | +1 | 6e-67 | 232 | 110/173 (64%) | poly(A) binding protein 2 | chr4:16336732-16339892 FORWARD LENGTH=629 |
