Probe CUST_33501_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_33501_PI426222305 | JHI_St_60k_v1 | DMT400045471 | CTAAGTACTACTAGTTTGACTATATGAGGTGCCTTTTGACTCTTTATTAGCTTTGTTCCA |
All Microarray Probes Designed to Gene DMG400017640
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_33501_PI426222305 | JHI_St_60k_v1 | DMT400045471 | CTAAGTACTACTAGTTTGACTATATGAGGTGCCTTTTGACTCTTTATTAGCTTTGTTCCA |
| CUST_33571_PI426222305 | JHI_St_60k_v1 | DMT400045470 | CTAAGTACTACTAGTTTGACTATATGAGGTGCCTTTTGACTCTTTATTAGCTTTGTTCCA |
Microarray Signals from CUST_33501_PI426222305
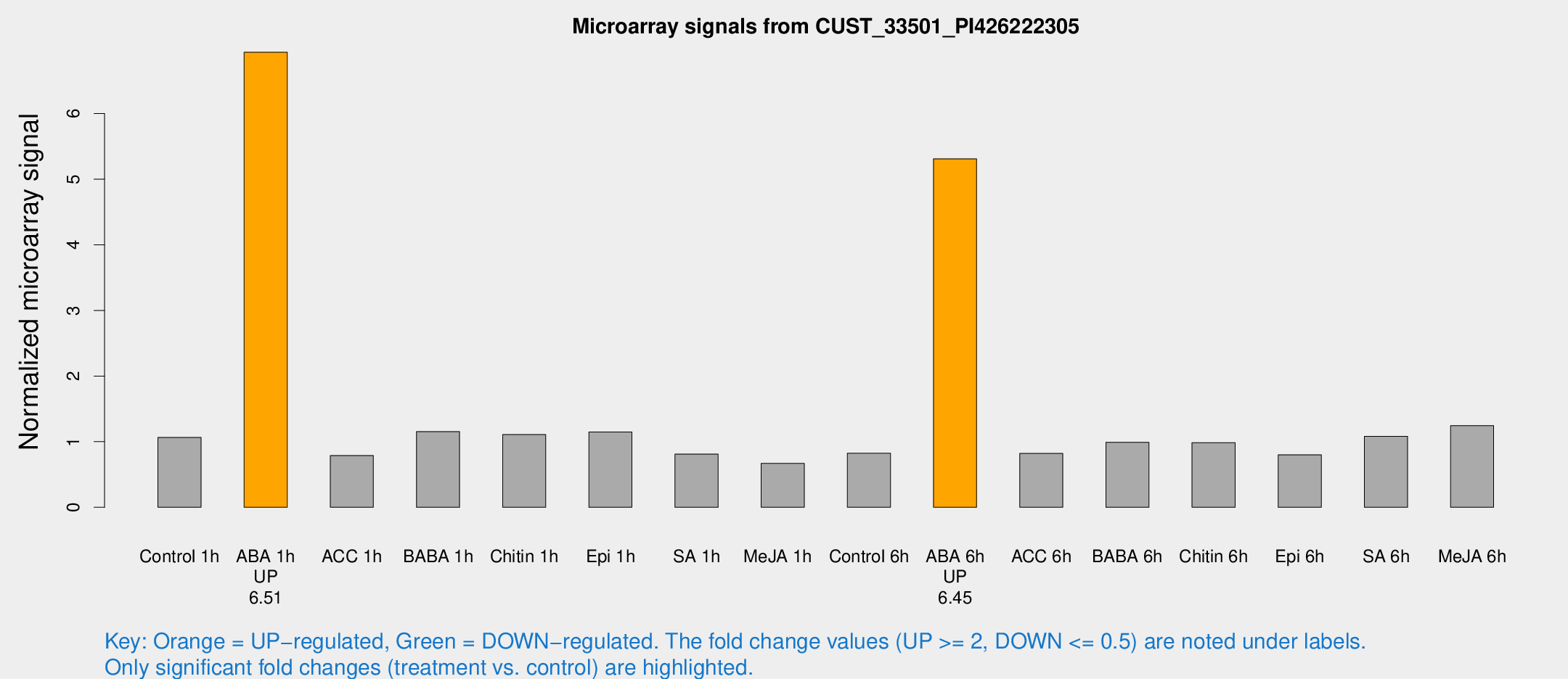
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 749.109 | 83.941 | 1.06463 | 0.0616181 |
| ABA 1h | 4388.12 | 695.174 | 6.93391 | 0.894495 |
| ACC 1h | 585.42 | 106.158 | 0.788892 | 0.0995756 |
| BABA 1h | 820.252 | 178.909 | 1.1544 | 0.174192 |
| Chitin 1h | 715.264 | 132.579 | 1.10818 | 0.114752 |
| Epi 1h | 701.374 | 84.6251 | 1.14882 | 0.112901 |
| SA 1h | 613.611 | 150.128 | 0.810821 | 0.147629 |
| Me-JA 1h | 388.2 | 59.3663 | 0.668885 | 0.0434912 |
| Control 6h | 603.296 | 122.62 | 0.823354 | 0.116661 |
| ABA 6h | 3986.97 | 492.866 | 5.31041 | 0.386943 |
| ACC 6h | 657.928 | 48.3893 | 0.820395 | 0.0857723 |
| BABA 6h | 779.758 | 83.2486 | 0.990623 | 0.0678257 |
| Chitin 6h | 729.608 | 42.2674 | 0.985907 | 0.0571061 |
| Epi 6h | 630.689 | 46.1949 | 0.800245 | 0.046421 |
| SA 6h | 749.376 | 64.5989 | 1.08162 | 0.0626343 |
| Me-JA 6h | 908.145 | 213.492 | 1.2446 | 0.186444 |
Source Transcript PGSC0003DMT400045471 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G01470.1 | +1 | 6e-44 | 158 | 97/244 (40%) | homeobox 1 | chr3:182648-184034 REVERSE LENGTH=272 |
