Probe CUST_33414_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_33414_PI426222305 | JHI_St_60k_v1 | DMT400013893 | AACACAATTAACATGGCTTCAAAACCAGGGATTCTGACCGAATGGCCATGGACATGGCTT |
All Microarray Probes Designed to Gene DMG400005436
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_33433_PI426222305 | JHI_St_60k_v1 | DMT400013894 | AACACAATTAACATGGCTTCAAAACCAGGGATTCTGACCGAATGGCCATGGACATGGCTT |
| CUST_33414_PI426222305 | JHI_St_60k_v1 | DMT400013893 | AACACAATTAACATGGCTTCAAAACCAGGGATTCTGACCGAATGGCCATGGACATGGCTT |
Microarray Signals from CUST_33414_PI426222305
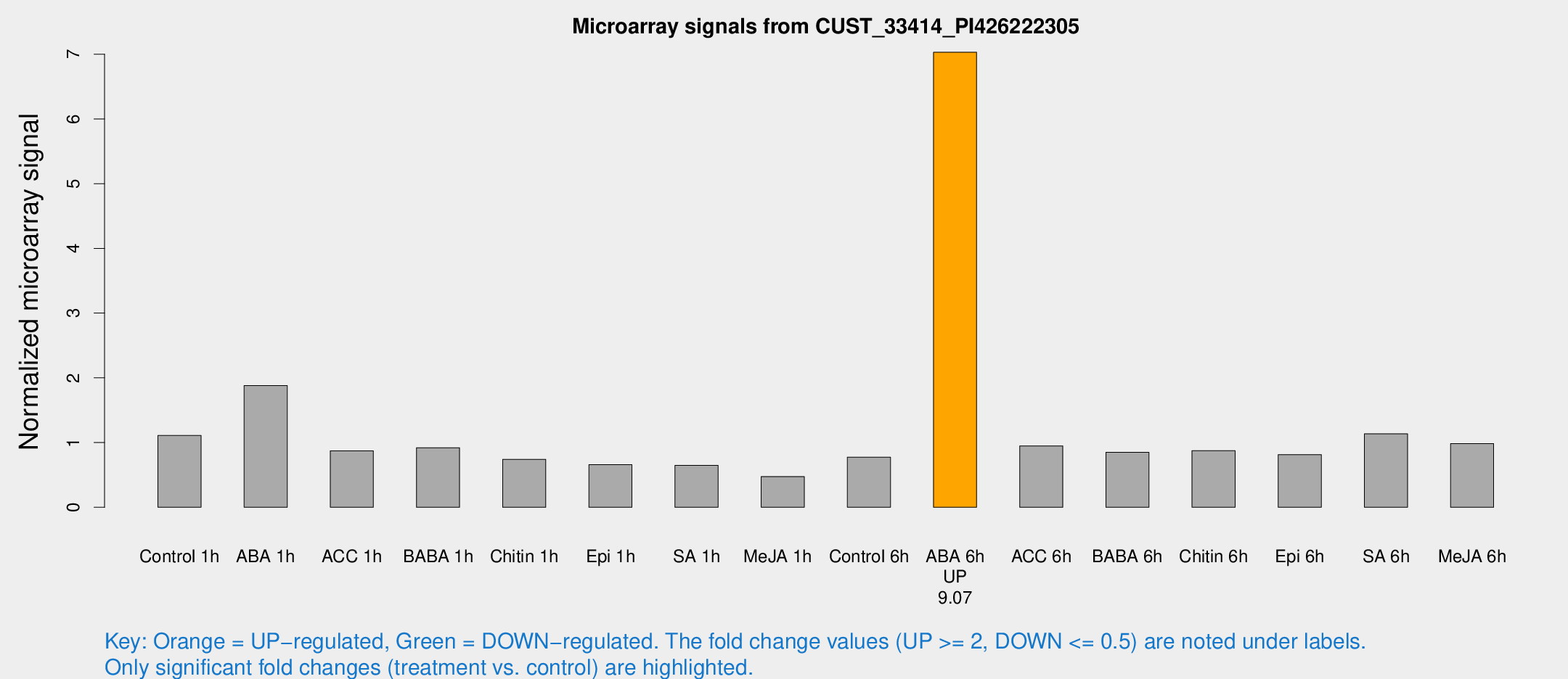
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 73.3219 | 8.21201 | 1.11027 | 0.084812 |
| ABA 1h | 110.018 | 12.692 | 1.88168 | 0.124709 |
| ACC 1h | 68.2574 | 22.0618 | 0.873294 | 0.337028 |
| BABA 1h | 59.498 | 10.3546 | 0.920547 | 0.110708 |
| Chitin 1h | 47.0912 | 13.8474 | 0.741075 | 0.247108 |
| Epi 1h | 39.6734 | 10.3682 | 0.658001 | 0.160945 |
| SA 1h | 43.5589 | 4.42197 | 0.646735 | 0.0662517 |
| Me-JA 1h | 27.916 | 9.26751 | 0.475058 | 0.20229 |
| Control 6h | 64.8527 | 25.099 | 0.775385 | 0.430107 |
| ABA 6h | 504.444 | 88.67 | 7.02999 | 1.01191 |
| ACC 6h | 72.3735 | 10.0328 | 0.949029 | 0.0979254 |
| BABA 6h | 66.0102 | 17.2214 | 0.849892 | 0.204984 |
| Chitin 6h | 62.4669 | 10.1418 | 0.875612 | 0.172532 |
| Epi 6h | 61.4368 | 10.3647 | 0.814329 | 0.13447 |
| SA 6h | 78.3721 | 19.2438 | 1.13456 | 0.471142 |
| Me-JA 6h | 72.0458 | 21.4532 | 0.984098 | 0.304826 |
Source Transcript PGSC0003DMT400013893 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G02205.3 | +3 | 0.0 | 523 | 259/372 (70%) | Fatty acid hydroxylase superfamily | chr1:418818-422154 FORWARD LENGTH=630 |
