Probe CUST_33248_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_33248_PI426222305 | JHI_St_60k_v1 | DMT400017915 | GTTGTTGTAATAATGACTCTGTTCTGCAGAGGGAATTGGAATAAATAATATTGGAGCAAC |
All Microarray Probes Designed to Gene DMG400006956
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_33248_PI426222305 | JHI_St_60k_v1 | DMT400017915 | GTTGTTGTAATAATGACTCTGTTCTGCAGAGGGAATTGGAATAAATAATATTGGAGCAAC |
| CUST_33300_PI426222305 | JHI_St_60k_v1 | DMT400017914 | GTTGTTGTAATAATGACTCTGTTCTGCAGAGGGAATTGGAATAAATAATATTGGAGCAAC |
Microarray Signals from CUST_33248_PI426222305
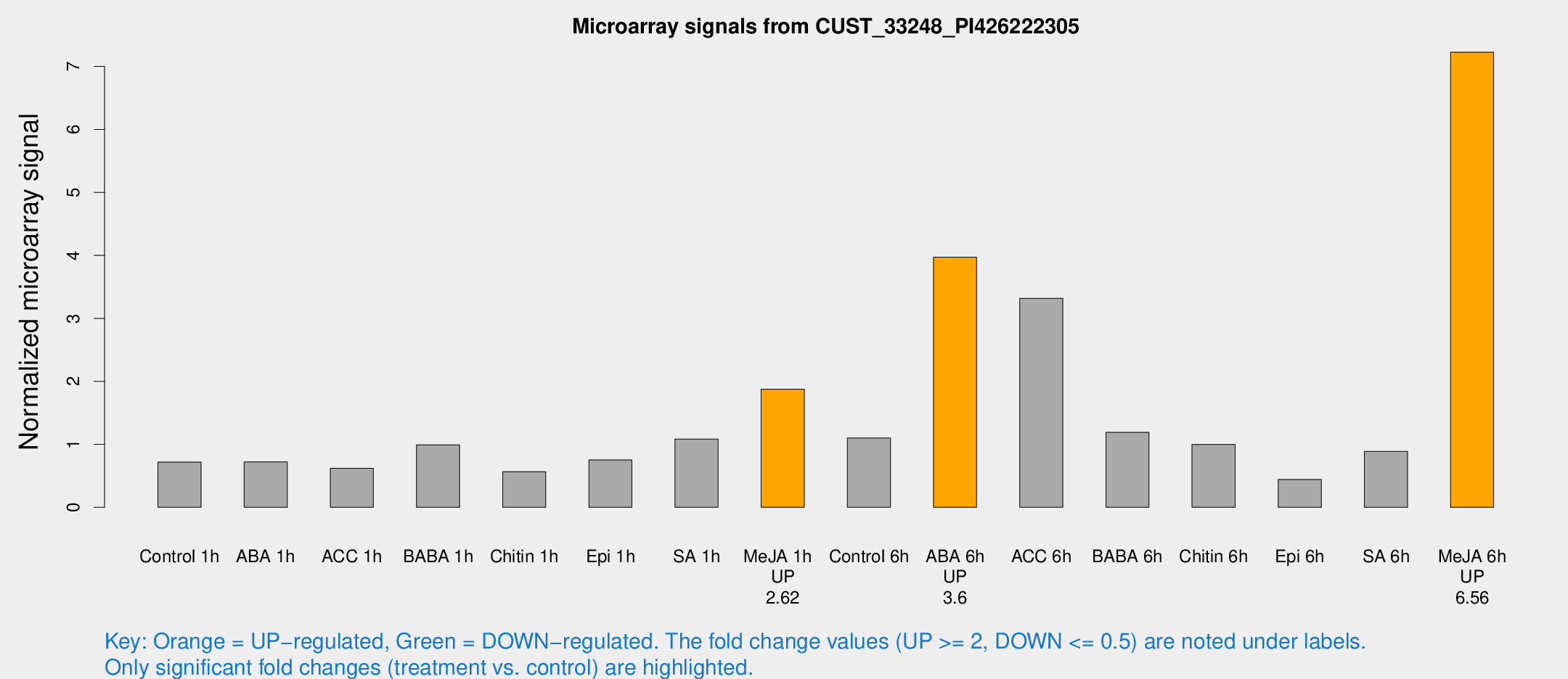
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5607.23 | 1019.36 | 0.716951 | 0.167976 |
| ABA 1h | 5249.74 | 1339.24 | 0.721868 | 0.193257 |
| ACC 1h | 6873.04 | 3025.88 | 0.621057 | 0.510028 |
| BABA 1h | 7391.1 | 992.041 | 0.992137 | 0.0572836 |
| Chitin 1h | 4117.74 | 943.558 | 0.566706 | 0.155445 |
| Epi 1h | 5458.46 | 1690.08 | 0.752171 | 0.258271 |
| SA 1h | 9678.75 | 3653.58 | 1.08288 | 0.473885 |
| Me-JA 1h | 11819.9 | 1551.04 | 1.87619 | 0.108323 |
| Control 6h | 9556.39 | 3650.98 | 1.10193 | 0.345556 |
| ABA 6h | 32603.6 | 4203.72 | 3.97154 | 0.549698 |
| ACC 6h | 32062 | 10209.2 | 3.3188 | 1.39487 |
| BABA 6h | 10924.9 | 3191.86 | 1.19114 | 0.296102 |
| Chitin 6h | 9719.14 | 3936.62 | 0.997992 | 0.555622 |
| Epi 6h | 4171.95 | 1153.58 | 0.442709 | 0.201776 |
| SA 6h | 6868.29 | 1263.25 | 0.887624 | 0.128658 |
| Me-JA 6h | 55227.8 | 5849.85 | 7.22596 | 1.13451 |
Source Transcript PGSC0003DMT400017915 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G01500.2 | +3 | 2e-61 | 199 | 95/137 (69%) | carbonic anhydrase 1 | chr3:194853-197873 REVERSE LENGTH=347 |
