Probe CUST_33148_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_33148_PI426222305 | JHI_St_60k_v1 | DMT400024622 | CATCAGATGTACCTTCTTTTGTTTTTAGGCCTGAAGCAGAAAGAATAATTGAAATGATGG |
All Microarray Probes Designed to Gene DMG400009527
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_33148_PI426222305 | JHI_St_60k_v1 | DMT400024622 | CATCAGATGTACCTTCTTTTGTTTTTAGGCCTGAAGCAGAAAGAATAATTGAAATGATGG |
Microarray Signals from CUST_33148_PI426222305
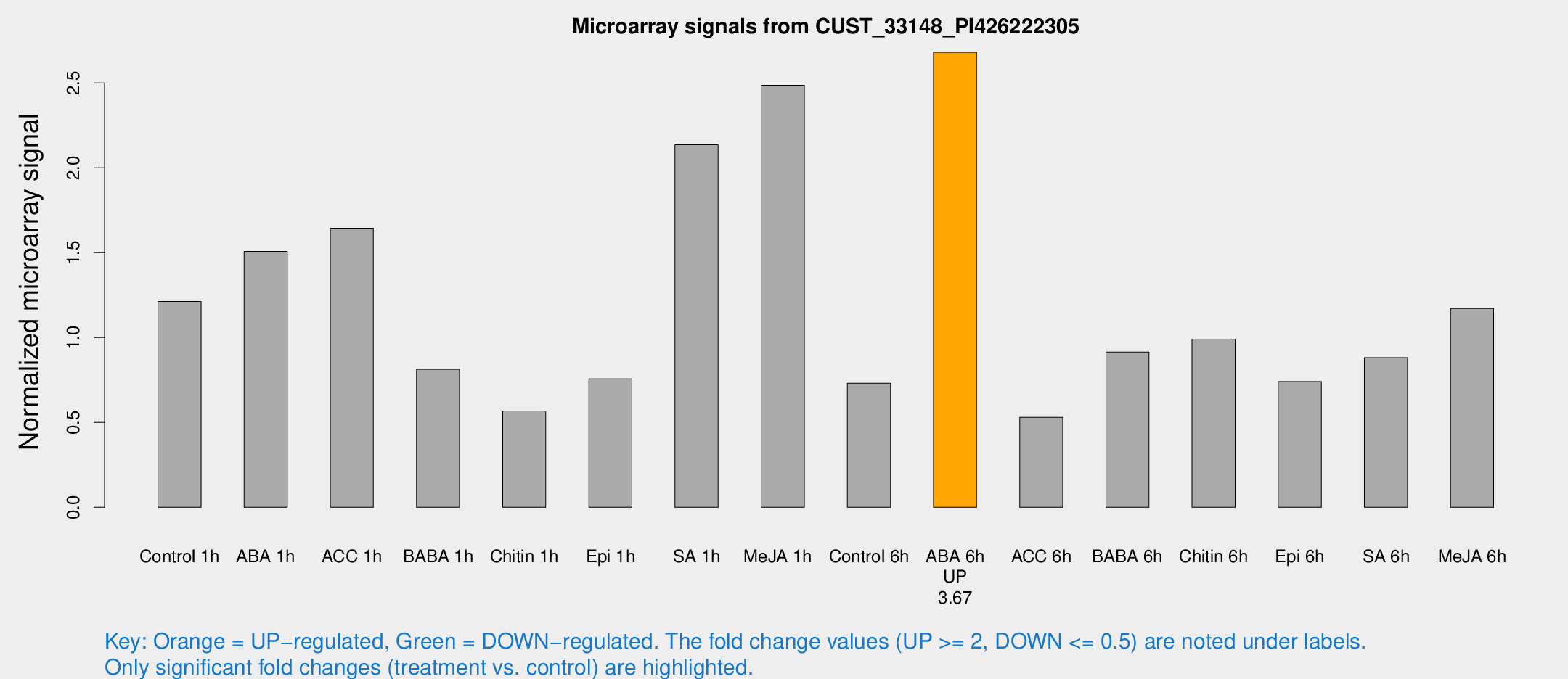
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 21.5242 | 4.77813 | 1.21238 | 0.214168 |
| ABA 1h | 30.8939 | 13.0433 | 1.50734 | 1.13463 |
| ACC 1h | 37.7861 | 14.8981 | 1.64414 | 1.07393 |
| BABA 1h | 15.1504 | 4.866 | 0.813166 | 0.246249 |
| Chitin 1h | 9.25332 | 2.97007 | 0.56765 | 0.22083 |
| Epi 1h | 11.7244 | 2.97309 | 0.756856 | 0.215058 |
| SA 1h | 37.9588 | 5.35472 | 2.13535 | 0.251509 |
| Me-JA 1h | 34.7958 | 3.65685 | 2.48549 | 0.49939 |
| Control 6h | 12.8907 | 3.09201 | 0.731163 | 0.195656 |
| ABA 6h | 49.3434 | 7.21687 | 2.68042 | 0.247586 |
| ACC 6h | 12.1094 | 4.97811 | 0.529798 | 0.199366 |
| BABA 6h | 18.1465 | 3.98871 | 0.914674 | 0.201766 |
| Chitin 6h | 17.9231 | 3.50898 | 0.990537 | 0.19497 |
| Epi 6h | 14.5126 | 3.61946 | 0.740943 | 0.202157 |
| SA 6h | 17.7073 | 5.98522 | 0.881135 | 0.341403 |
| Me-JA 6h | 20.4689 | 3.73268 | 1.17118 | 0.200063 |
Source Transcript PGSC0003DMT400024622 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G05675.1 | +1 | 9e-143 | 420 | 217/448 (48%) | UDP-Glycosyltransferase superfamily protein | chr1:1701213-1702715 REVERSE LENGTH=453 |
