Probe CUST_33141_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_33141_PI426222305 | JHI_St_60k_v1 | DMT400067578 | GAGTTTGTTGGAGGTGACATGTTTCAAAAGATTCCTAATGCTAATGCAATCTTACTCAAG |
All Microarray Probes Designed to Gene DMG400026275
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_33141_PI426222305 | JHI_St_60k_v1 | DMT400067578 | GAGTTTGTTGGAGGTGACATGTTTCAAAAGATTCCTAATGCTAATGCAATCTTACTCAAG |
| CUST_33224_PI426222305 | JHI_St_60k_v1 | DMT400067579 | AACTTAATCATATGTGTCTTCGTATTTTGCATTGGGTTTGAGTATGTACGTGTGAACAAC |
Microarray Signals from CUST_33141_PI426222305
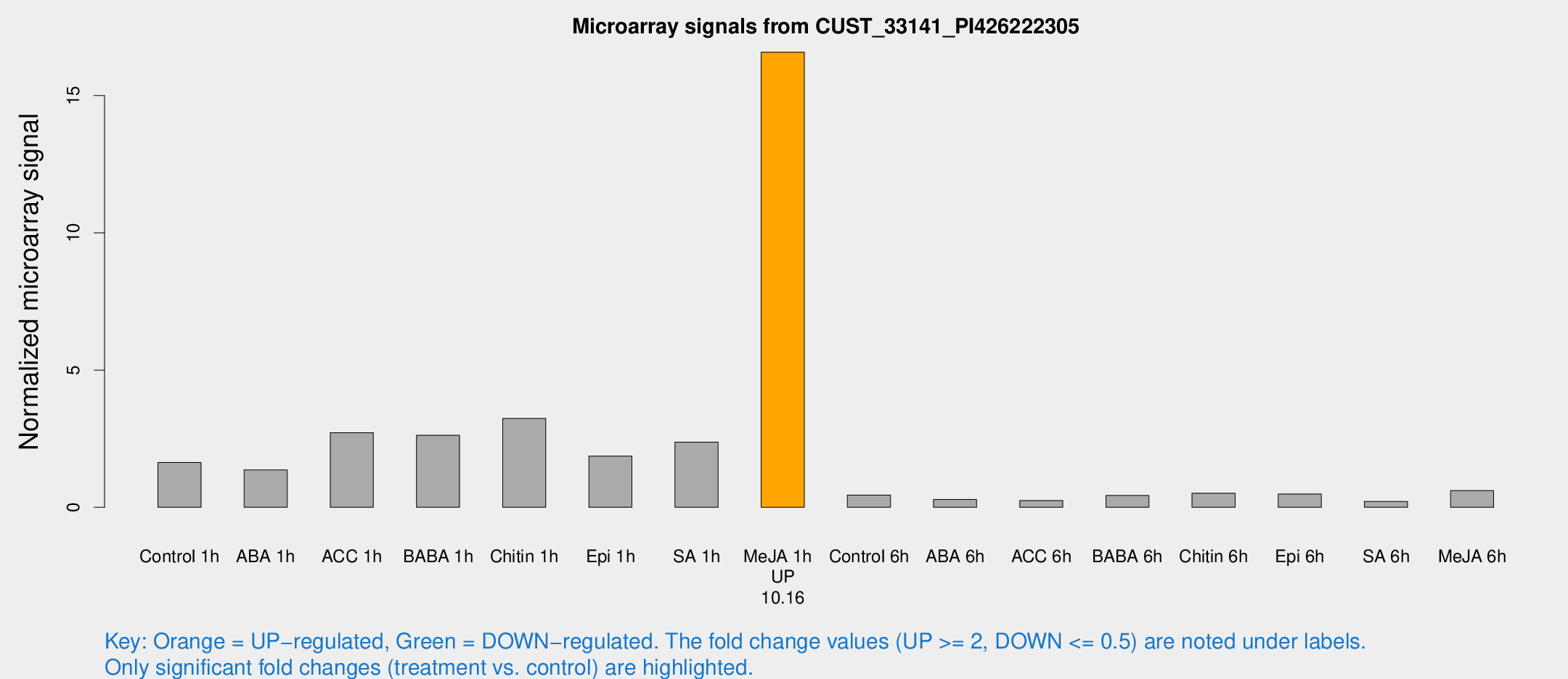
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 50.9938 | 9.59791 | 1.63119 | 0.280506 |
| ABA 1h | 45.9768 | 22.2866 | 1.36319 | 0.690944 |
| ACC 1h | 84.9279 | 8.23107 | 2.72071 | 0.481667 |
| BABA 1h | 80.9498 | 18.0747 | 2.62351 | 0.409216 |
| Chitin 1h | 97.3281 | 33.0221 | 3.2391 | 1.12567 |
| Epi 1h | 49.5066 | 6.36783 | 1.86453 | 0.311273 |
| SA 1h | 75.206 | 12.0217 | 2.37161 | 0.226993 |
| Me-JA 1h | 408.768 | 23.901 | 16.5802 | 1.33305 |
| Control 6h | 15.2157 | 4.63257 | 0.44684 | 0.153462 |
| ABA 6h | 10.2829 | 3.76682 | 0.285242 | 0.133524 |
| ACC 6h | 8.61719 | 4.31053 | 0.245058 | 0.123595 |
| BABA 6h | 15.7579 | 4.8341 | 0.429252 | 0.12619 |
| Chitin 6h | 18.3145 | 5.06892 | 0.513722 | 0.180062 |
| Epi 6h | 18.2341 | 4.92143 | 0.486 | 0.222285 |
| SA 6h | 6.46267 | 3.75013 | 0.216668 | 0.125618 |
| Me-JA 6h | 23.6402 | 8.93321 | 0.612909 | 0.362993 |
Source Transcript PGSC0003DMT400067578 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G35160.1 | +3 | 2e-56 | 194 | 131/365 (36%) | O-methyltransferase family protein | chr4:16730989-16732808 REVERSE LENGTH=382 |
