Probe CUST_32882_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_32882_PI426222305 | JHI_St_60k_v1 | DMT400062617 | CCAAAGATTCAGAAATCATATACACATGGCTAGTGTTGAAGCAAGACATCTCAATCTTTT |
All Microarray Probes Designed to Gene DMG400024368
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_32882_PI426222305 | JHI_St_60k_v1 | DMT400062617 | CCAAAGATTCAGAAATCATATACACATGGCTAGTGTTGAAGCAAGACATCTCAATCTTTT |
| CUST_32865_PI426222305 | JHI_St_60k_v1 | DMT400062618 | GGATATCAATTCCATCATTTCTCACCATGTAAGCTTAATCCCCAATTCAATTCTCTCTGT |
Microarray Signals from CUST_32882_PI426222305
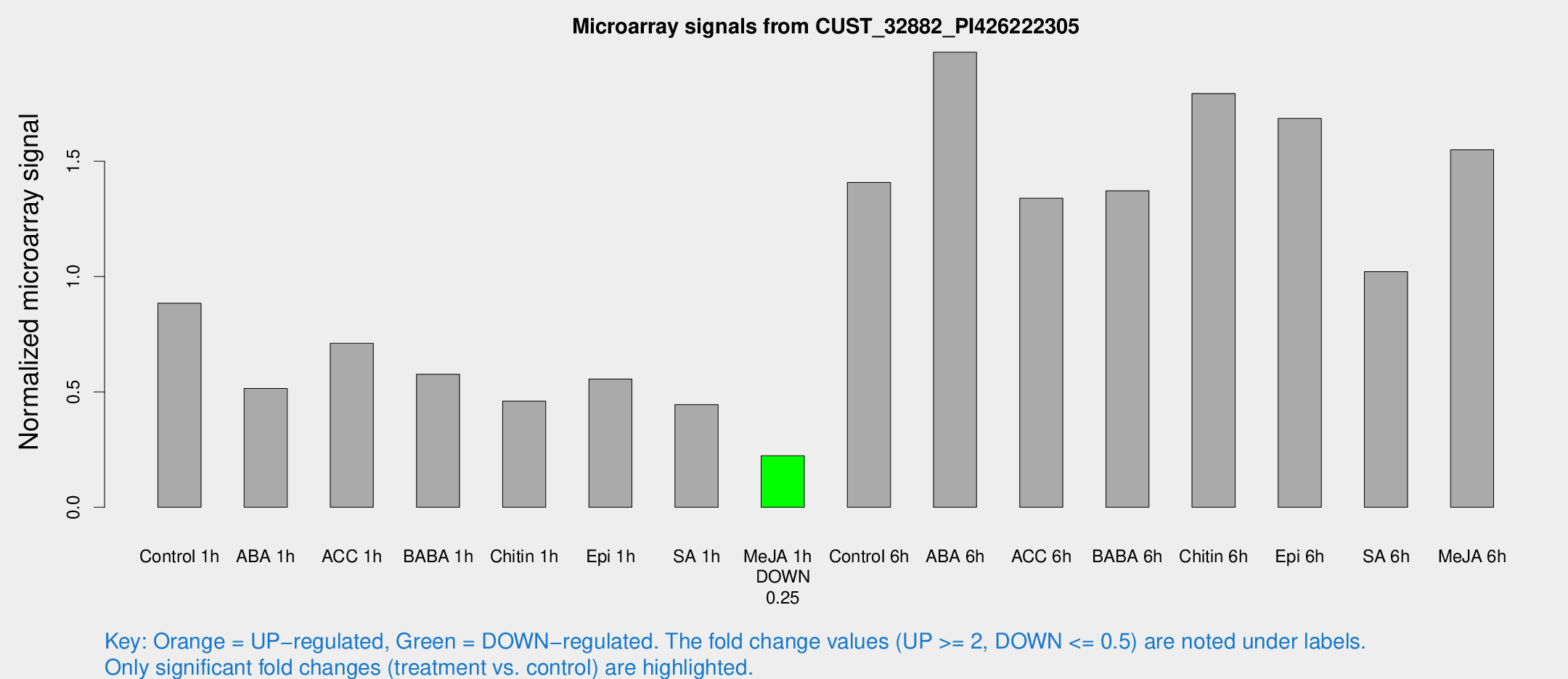
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 46.9233 | 5.19023 | 0.884723 | 0.0751735 |
| ABA 1h | 24.2116 | 3.1014 | 0.514641 | 0.0683801 |
| ACC 1h | 43.5823 | 13.3448 | 0.710596 | 0.241242 |
| BABA 1h | 29.4196 | 3.47289 | 0.576699 | 0.0694409 |
| Chitin 1h | 24.9586 | 9.34211 | 0.459661 | 0.169637 |
| Epi 1h | 25.4239 | 3.20265 | 0.556255 | 0.072018 |
| SA 1h | 25.3292 | 6.06514 | 0.445191 | 0.0938149 |
| Me-JA 1h | 9.5813 | 2.99132 | 0.223141 | 0.0695581 |
| Control 6h | 78.414 | 17.7732 | 1.40782 | 0.244022 |
| ABA 6h | 116.072 | 25.4428 | 1.9721 | 0.431275 |
| ACC 6h | 84.5225 | 19.3586 | 1.33983 | 0.106755 |
| BABA 6h | 84.1297 | 18.7581 | 1.37135 | 0.284868 |
| Chitin 6h | 102.487 | 15.2355 | 1.79298 | 0.308734 |
| Epi 6h | 99.7491 | 6.71384 | 1.68534 | 0.112965 |
| SA 6h | 57.1976 | 14.1331 | 1.02121 | 0.183967 |
| Me-JA 6h | 88.3613 | 27.2248 | 1.54941 | 0.344812 |
Source Transcript PGSC0003DMT400062617 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G12920.1 | +3 | 2e-75 | 240 | 142/316 (45%) | SBP (S-ribonuclease binding protein) family protein | chr3:4122127-4123323 REVERSE LENGTH=335 |
