Probe CUST_32724_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_32724_PI426222305 | JHI_St_60k_v1 | DMT400047290 | ATTGACTCCTTGAGCCGTGAGTAATCTGTTACACTTTCTACCAAAGGCGACTTCATTTCC |
All Microarray Probes Designed to Gene DMG400018364
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_32724_PI426222305 | JHI_St_60k_v1 | DMT400047290 | ATTGACTCCTTGAGCCGTGAGTAATCTGTTACACTTTCTACCAAAGGCGACTTCATTTCC |
| CUST_32683_PI426222305 | JHI_St_60k_v1 | DMT400047289 | TTGAGTTGTTTGAATCAAGAACAGAGTCAAATTGTCCCTGTAAATTGTTCTGTTAAACCC |
Microarray Signals from CUST_32724_PI426222305
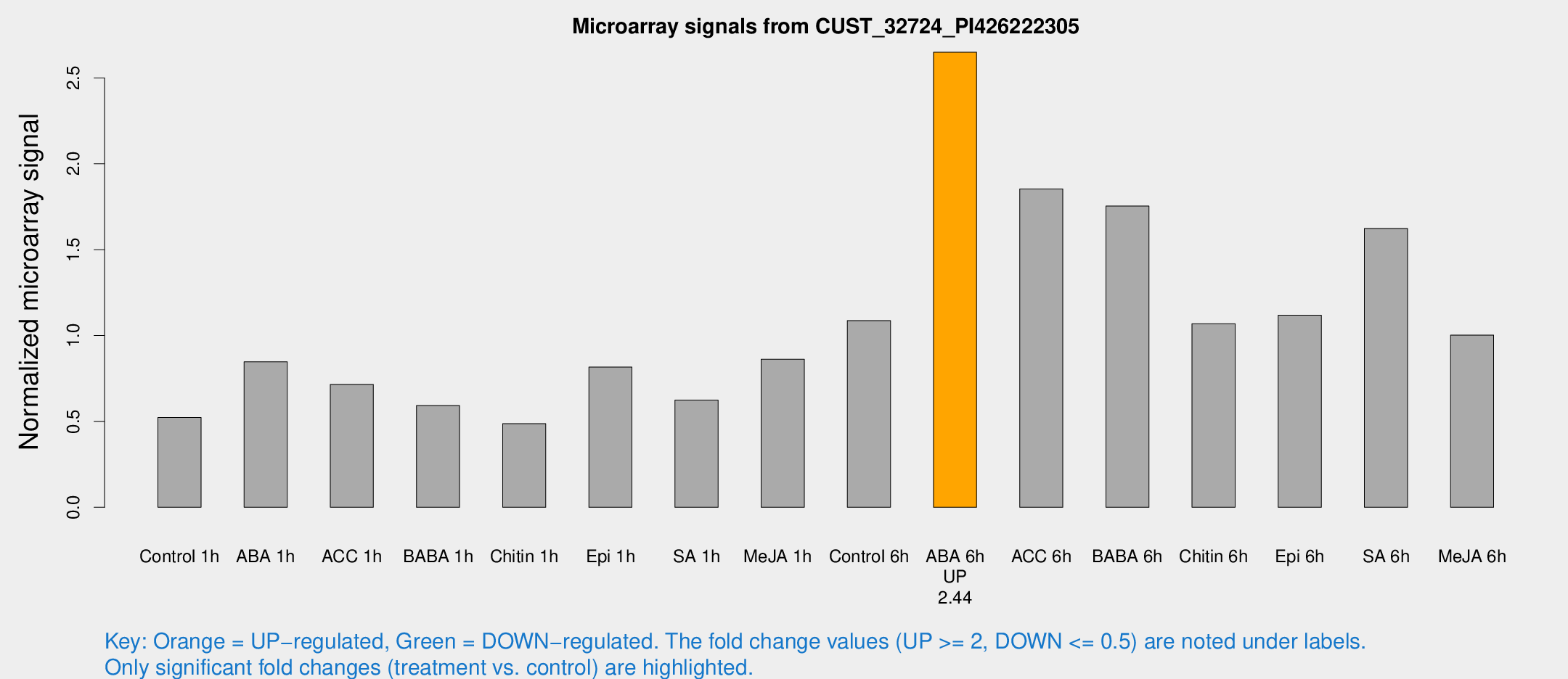
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 15.4506 | 4.63393 | 0.522788 | 0.152557 |
| ABA 1h | 24.4511 | 9.96416 | 0.847333 | 0.392862 |
| ACC 1h | 21.3632 | 5.58086 | 0.715547 | 0.257553 |
| BABA 1h | 18.2093 | 5.8814 | 0.592376 | 0.212239 |
| Chitin 1h | 13.0441 | 3.66736 | 0.486896 | 0.176916 |
| Epi 1h | 19.5297 | 3.64051 | 0.816215 | 0.15677 |
| SA 1h | 20.8402 | 8.00951 | 0.62437 | 0.286621 |
| Me-JA 1h | 19.4736 | 3.87031 | 0.861895 | 0.175535 |
| Control 6h | 29.871 | 4.04166 | 1.08732 | 0.149577 |
| ABA 6h | 77.3179 | 6.94675 | 2.64953 | 0.201342 |
| ACC 6h | 59.2738 | 8.82763 | 1.85385 | 0.20662 |
| BABA 6h | 54.4957 | 7.44609 | 1.75403 | 0.173986 |
| Chitin 6h | 32.3465 | 6.49697 | 1.06845 | 0.272327 |
| Epi 6h | 34.4792 | 4.82223 | 1.11842 | 0.156053 |
| SA 6h | 43.9921 | 4.57126 | 1.62354 | 0.170666 |
| Me-JA 6h | 36.0517 | 14.3227 | 1.0025 | 0.672383 |
Source Transcript PGSC0003DMT400047290 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G03530.1 | +3 | 1e-53 | 202 | 155/364 (43%) | nuclear assembly factor 1 | chr1:880638-883338 REVERSE LENGTH=801 |
