Probe CUST_31507_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_31507_PI426222305 | JHI_St_60k_v1 | DMT400042641 | CAGCCATTATAGGTGTTGAGATAGCCTTCTTGATCTCTTCAATTAGACATACGTCCGCCA |
All Microarray Probes Designed to Gene DMG400016535
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_31507_PI426222305 | JHI_St_60k_v1 | DMT400042641 | CAGCCATTATAGGTGTTGAGATAGCCTTCTTGATCTCTTCAATTAGACATACGTCCGCCA |
| CUST_31515_PI426222305 | JHI_St_60k_v1 | DMT400042642 | CAGCCATTATAGGTGTTGAGATAGCCTTCTTGATCTCTTCAATTAGACATACGTCCGCCA |
Microarray Signals from CUST_31507_PI426222305
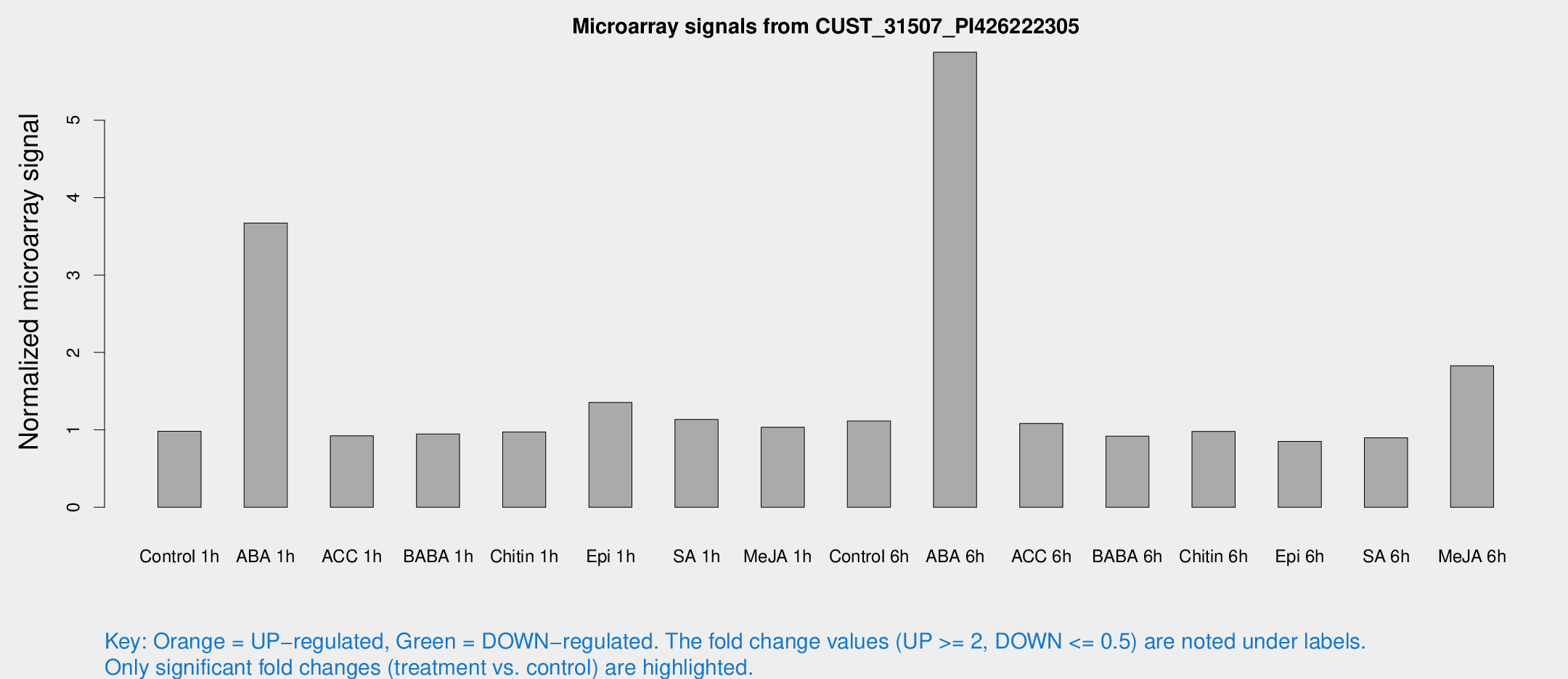
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 6.77623 | 3.09582 | 0.982444 | 0.487993 |
| ABA 1h | 24.1822 | 8.69056 | 3.67138 | 1.34419 |
| ACC 1h | 6.29036 | 3.36656 | 0.923733 | 0.494442 |
| BABA 1h | 6.07584 | 3.29364 | 0.94609 | 0.516866 |
| Chitin 1h | 5.7886 | 3.14836 | 0.973191 | 0.526203 |
| Epi 1h | 8.12563 | 3.15448 | 1.35455 | 0.583226 |
| SA 1h | 8.42549 | 3.15035 | 1.13477 | 0.506316 |
| Me-JA 1h | 5.6096 | 3.20059 | 1.03315 | 0.589531 |
| Control 6h | 7.59139 | 3.27673 | 1.11475 | 0.506396 |
| ABA 6h | 44.8245 | 11.4227 | 5.87759 | 1.61305 |
| ACC 6h | 9.22939 | 4.00353 | 1.08312 | 0.515923 |
| BABA 6h | 6.92559 | 3.66393 | 0.918263 | 0.494781 |
| Chitin 6h | 6.99903 | 3.57695 | 0.98093 | 0.514473 |
| Epi 6h | 6.40399 | 3.73684 | 0.850779 | 0.492649 |
| SA 6h | 5.8935 | 3.41528 | 0.898629 | 0.520393 |
| Me-JA 6h | 13.5198 | 3.85177 | 1.82825 | 0.787248 |
Source Transcript PGSC0003DMT400042641 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G79900.1 | +2 | 7e-107 | 321 | 157/227 (69%) | Mitochondrial substrate carrier family protein | chr1:30052524-30053599 REVERSE LENGTH=296 |
