Probe CUST_3126_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_3126_PI426222305 | JHI_St_60k_v1 | DMT400000245 | GAAATCATAATGAGTCTTCTGCTGAAACTAGACGCTATTCAGGTATGTGGATAACATTAA |
All Microarray Probes Designed to Gene DMG400000079
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_3126_PI426222305 | JHI_St_60k_v1 | DMT400000245 | GAAATCATAATGAGTCTTCTGCTGAAACTAGACGCTATTCAGGTATGTGGATAACATTAA |
| CUST_2978_PI426222305 | JHI_St_60k_v1 | DMT400000247 | TTTGGCACTGAGGCGTGACAATATCGACTTCAAAGTTACTTGTTTGGTATTGAGGCGTGA |
Microarray Signals from CUST_3126_PI426222305
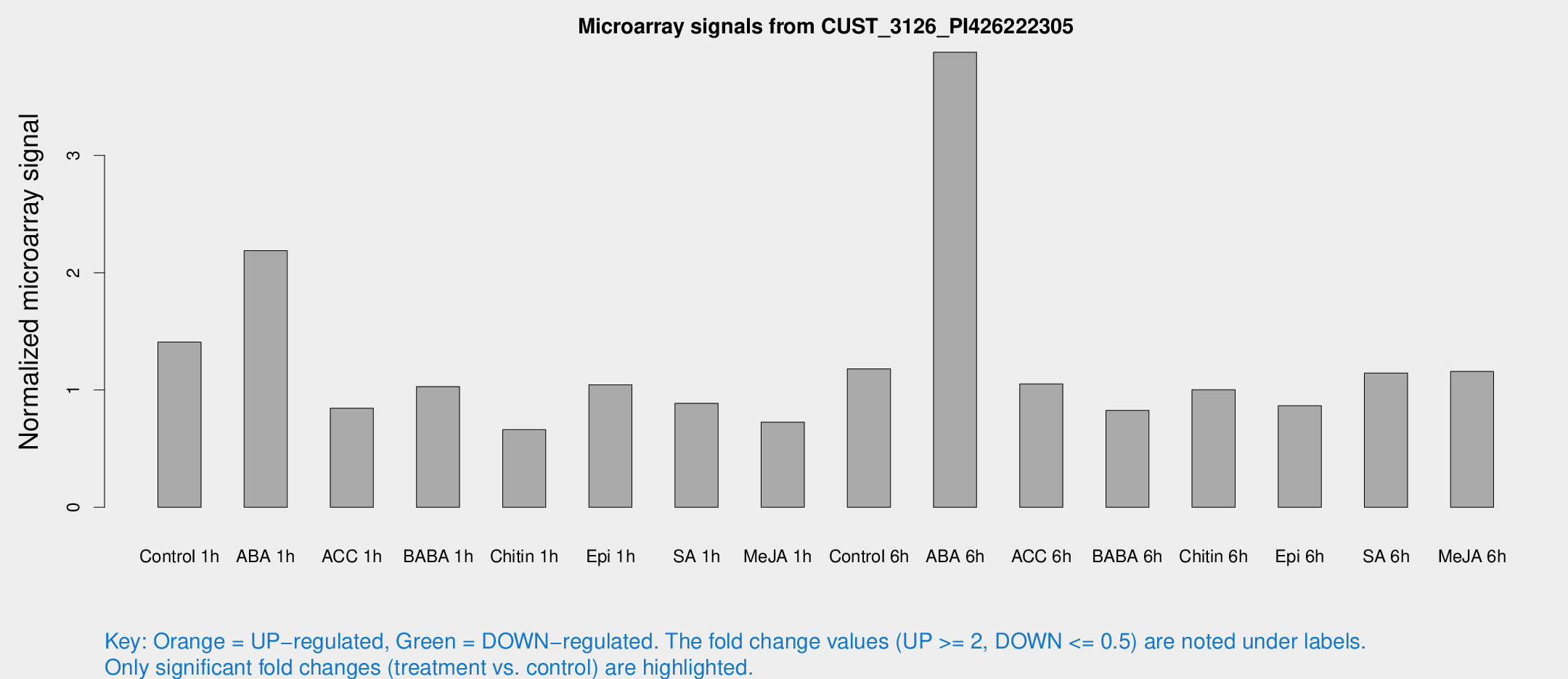
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 520.675 | 70.9604 | 1.40931 | 0.248161 |
| ABA 1h | 854.549 | 395.595 | 2.18838 | 1.0968 |
| ACC 1h | 429.601 | 208.257 | 0.845309 | 0.563907 |
| BABA 1h | 378.765 | 82.0271 | 1.02908 | 0.143244 |
| Chitin 1h | 225.191 | 42.7831 | 0.662271 | 0.141504 |
| Epi 1h | 382.903 | 155.422 | 1.04475 | 0.454523 |
| SA 1h | 342.644 | 62.846 | 0.885917 | 0.122017 |
| Me-JA 1h | 217.507 | 23.9996 | 0.725349 | 0.0749688 |
| Control 6h | 440.529 | 74.0116 | 1.18004 | 0.165367 |
| ABA 6h | 1578.1 | 331.057 | 3.87935 | 0.80992 |
| ACC 6h | 460.445 | 106.396 | 1.05249 | 0.304204 |
| BABA 6h | 344.317 | 57.505 | 0.825549 | 0.122198 |
| Chitin 6h | 389.702 | 30.6546 | 1.00252 | 0.10872 |
| Epi 6h | 362.539 | 52.0087 | 0.8658 | 0.0777931 |
| SA 6h | 438.268 | 99.7168 | 1.14472 | 0.447214 |
| Me-JA 6h | 425.987 | 57.0484 | 1.15845 | 0.0675015 |
Source Transcript PGSC0003DMT400000245 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G46240.1 | +2 | 1e-21 | 100 | 99/384 (26%) | BCL-2-associated athanogene 6 | chr2:18986586-18989827 FORWARD LENGTH=1043 |
