Probe CUST_30951_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_30951_PI426222305 | JHI_St_60k_v1 | DMT400040283 | CTTGAATCCTTGACCTAAAATAAATTGAGGCTTCTTTCTTTAAGACAGAAAATGAGTGCC |
All Microarray Probes Designed to Gene DMG400015594
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_30963_PI426222305 | JHI_St_60k_v1 | DMT400040284 | GGCTATTGCAATTGGAACTATCTCCAGTCTTATTGAGTAAACAAGGAATTTGAACAACAA |
| CUST_30951_PI426222305 | JHI_St_60k_v1 | DMT400040283 | CTTGAATCCTTGACCTAAAATAAATTGAGGCTTCTTTCTTTAAGACAGAAAATGAGTGCC |
Microarray Signals from CUST_30951_PI426222305
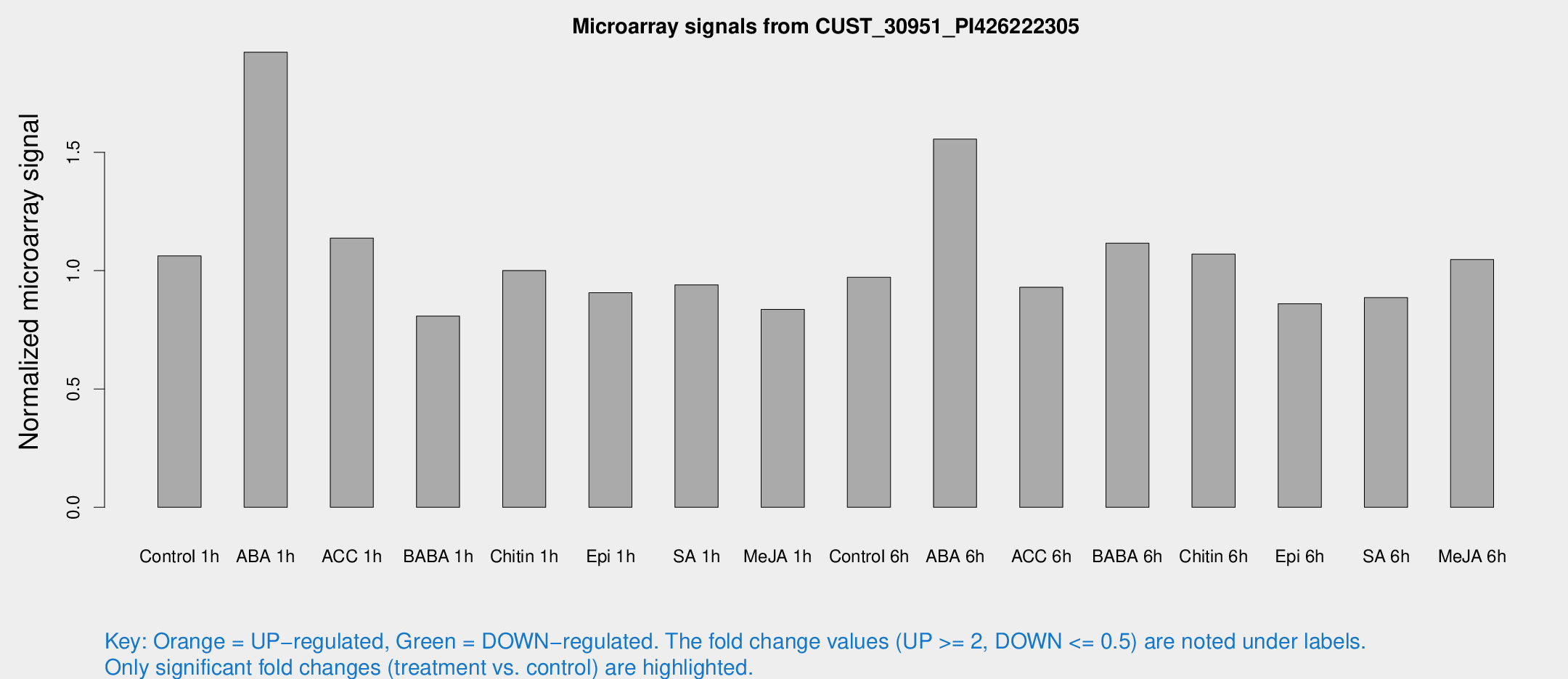
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 437.538 | 40.27 | 1.06182 | 0.0619708 |
| ABA 1h | 696.157 | 40.406 | 1.92254 | 0.111423 |
| ACC 1h | 479.137 | 28.4823 | 1.13737 | 0.111878 |
| BABA 1h | 324.126 | 41.9595 | 0.807423 | 0.0476926 |
| Chitin 1h | 372.888 | 42.7215 | 1.00022 | 0.0811227 |
| Epi 1h | 321.758 | 18.9767 | 0.906408 | 0.0532986 |
| SA 1h | 401.522 | 50.0934 | 0.939857 | 0.0817465 |
| Me-JA 1h | 279.763 | 16.6336 | 0.836376 | 0.0524159 |
| Control 6h | 412.39 | 73.4036 | 0.971563 | 0.0959439 |
| ABA 6h | 685.023 | 81.8917 | 1.55534 | 0.10829 |
| ACC 6h | 443.037 | 55.4015 | 0.92959 | 0.0611394 |
| BABA 6h | 511.05 | 29.842 | 1.1159 | 0.0650981 |
| Chitin 6h | 466.592 | 27.3455 | 1.06972 | 0.0808542 |
| Epi 6h | 403.013 | 50.1849 | 0.859929 | 0.182018 |
| SA 6h | 375.278 | 73.4186 | 0.886301 | 0.0937476 |
| Me-JA 6h | 450.793 | 105.853 | 1.04657 | 0.164877 |
Source Transcript PGSC0003DMT400040283 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G52190.1 | +2 | 1e-72 | 241 | 122/275 (44%) | Major facilitator superfamily protein | chr1:19434671-19438673 FORWARD LENGTH=607 |
