Probe CUST_30763_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_30763_PI426222305 | JHI_St_60k_v1 | DMT400022130 | CAAAAACATGATAGAAGAGAATCCTGGCAATTCCTTATTCTTGAGGAATTATGCCAACTT |
All Microarray Probes Designed to Gene DMG400008584
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_30758_PI426222305 | JHI_St_60k_v1 | DMT400022129 | CAAAAACATGATAGAAGAGAATCCTGGCAATTCCTTATTCTTGAGGAATTATGCCAACTT |
| CUST_30763_PI426222305 | JHI_St_60k_v1 | DMT400022130 | CAAAAACATGATAGAAGAGAATCCTGGCAATTCCTTATTCTTGAGGAATTATGCCAACTT |
Microarray Signals from CUST_30763_PI426222305
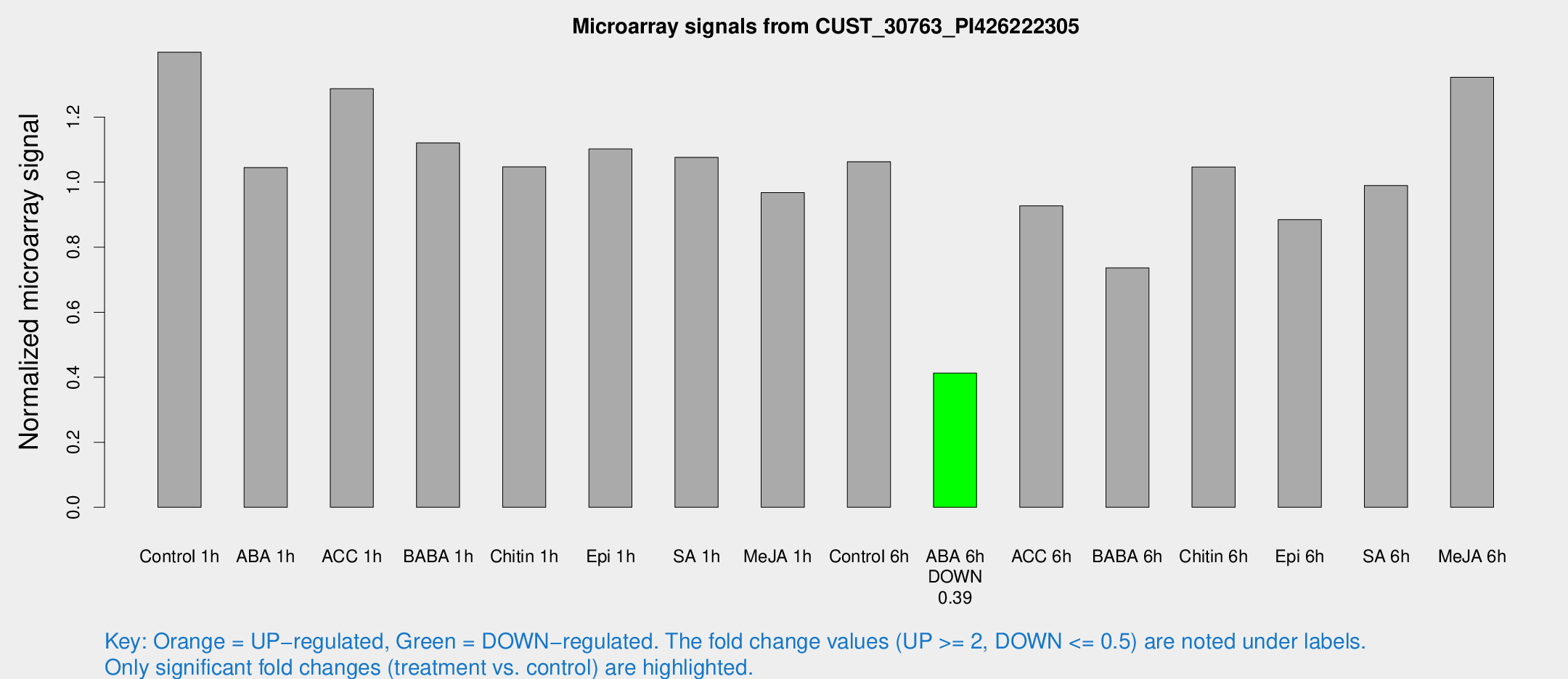
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 14557 | 1885.65 | 1.39957 | 0.086384 |
| ABA 1h | 9533.44 | 863.584 | 1.04477 | 0.111303 |
| ACC 1h | 14131.4 | 2725.73 | 1.28752 | 0.185166 |
| BABA 1h | 12042.4 | 3163.76 | 1.12068 | 0.236583 |
| Chitin 1h | 9875.08 | 1597.17 | 1.04704 | 0.0868758 |
| Epi 1h | 9836.82 | 868.806 | 1.10208 | 0.115757 |
| SA 1h | 12033.5 | 2678.1 | 1.07612 | 0.236012 |
| Me-JA 1h | 8270.05 | 1167.5 | 0.967702 | 0.0863444 |
| Control 6h | 11302.1 | 2106.57 | 1.06284 | 0.136057 |
| ABA 6h | 4500.33 | 260.317 | 0.412826 | 0.0238365 |
| ACC 6h | 10891.7 | 629.146 | 0.927217 | 0.108491 |
| BABA 6h | 8711.07 | 1562.19 | 0.736526 | 0.145009 |
| Chitin 6h | 11425.4 | 660.852 | 1.04674 | 0.0604342 |
| Epi 6h | 10327 | 1053.93 | 0.884814 | 0.0510858 |
| SA 6h | 10115.3 | 955.509 | 0.989796 | 0.0571469 |
| Me-JA 6h | 13806.8 | 2072.42 | 1.32286 | 0.211095 |
Source Transcript PGSC0003DMT400022130 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G20190.1 | +3 | 7e-27 | 110 | 50/94 (53%) | Tetratricopeptide repeat (TPR)-like superfamily protein | chr5:6814093-6815171 FORWARD LENGTH=290 |
