Probe CUST_30572_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_30572_PI426222305 | JHI_St_60k_v1 | DMT400025118 | CTGGATGATTTACAGGCAAAAACAAGTGTAATTATTGCCTCAGATAATCAGGATGATGAT |
All Microarray Probes Designed to Gene DMG400009705
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_30524_PI426222305 | JHI_St_60k_v1 | DMT400025119 | ATTTGAATTTCTACTGCTCTTTCGAGACAACAGTGAAGGCTGTGTGCAGGCCTGCGATTT |
| CUST_30572_PI426222305 | JHI_St_60k_v1 | DMT400025118 | CTGGATGATTTACAGGCAAAAACAAGTGTAATTATTGCCTCAGATAATCAGGATGATGAT |
Microarray Signals from CUST_30572_PI426222305
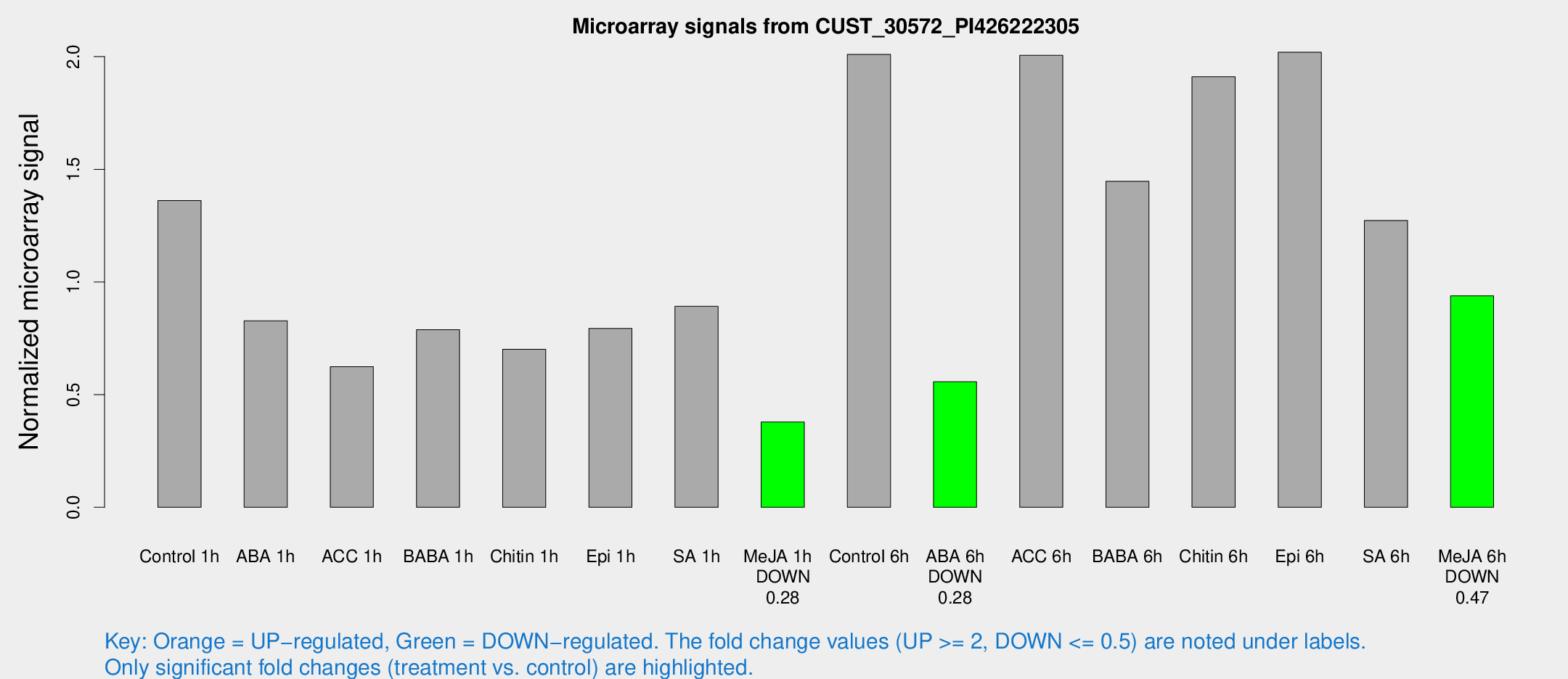
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1443.18 | 98.5501 | 1.36111 | 0.0786613 |
| ABA 1h | 775.592 | 51.8658 | 0.82726 | 0.0521281 |
| ACC 1h | 808.977 | 275.685 | 0.623925 | 0.27181 |
| BABA 1h | 906.153 | 284.88 | 0.788244 | 0.223471 |
| Chitin 1h | 668.178 | 40.4885 | 0.701638 | 0.0526021 |
| Epi 1h | 735.884 | 87.0735 | 0.793467 | 0.0882143 |
| SA 1h | 971.981 | 68.3042 | 0.892083 | 0.0926456 |
| Me-JA 1h | 334.849 | 57.3866 | 0.378311 | 0.0385038 |
| Control 6h | 2210.24 | 403.189 | 2.00958 | 0.232464 |
| ABA 6h | 632.916 | 70.7978 | 0.556844 | 0.0352538 |
| ACC 6h | 2447.81 | 220.393 | 2.00569 | 0.132187 |
| BABA 6h | 1747.32 | 246.681 | 1.44671 | 0.172788 |
| Chitin 6h | 2161.72 | 178.81 | 1.91077 | 0.161794 |
| Epi 6h | 2423.58 | 222.162 | 2.01952 | 0.136479 |
| SA 6h | 1487.6 | 424.897 | 1.27289 | 0.322156 |
| Me-JA 6h | 1027.33 | 195.983 | 0.938279 | 0.118458 |
Source Transcript PGSC0003DMT400025118 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G18210.1 | +1 | 2e-89 | 278 | 154/360 (43%) | purine permease 10 | chr4:10076175-10077495 FORWARD LENGTH=390 |
