Probe CUST_30376_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_30376_PI426222305 | JHI_St_60k_v1 | DMT400069751 | ATACACCGGAGGAATGGTGCCTGATGTAAACCAGGTATTTGCAAATATGTGATCAAGGGC |
All Microarray Probes Designed to Gene DMG400027125
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_30449_PI426222305 | JHI_St_60k_v1 | DMT400069750 | GGGAGATAGGTTAAAGACAACAAGGTCTTGTTAAGAAATGGGAATTGGAACATTTTTTGT |
| CUST_30376_PI426222305 | JHI_St_60k_v1 | DMT400069751 | ATACACCGGAGGAATGGTGCCTGATGTAAACCAGGTATTTGCAAATATGTGATCAAGGGC |
Microarray Signals from CUST_30376_PI426222305
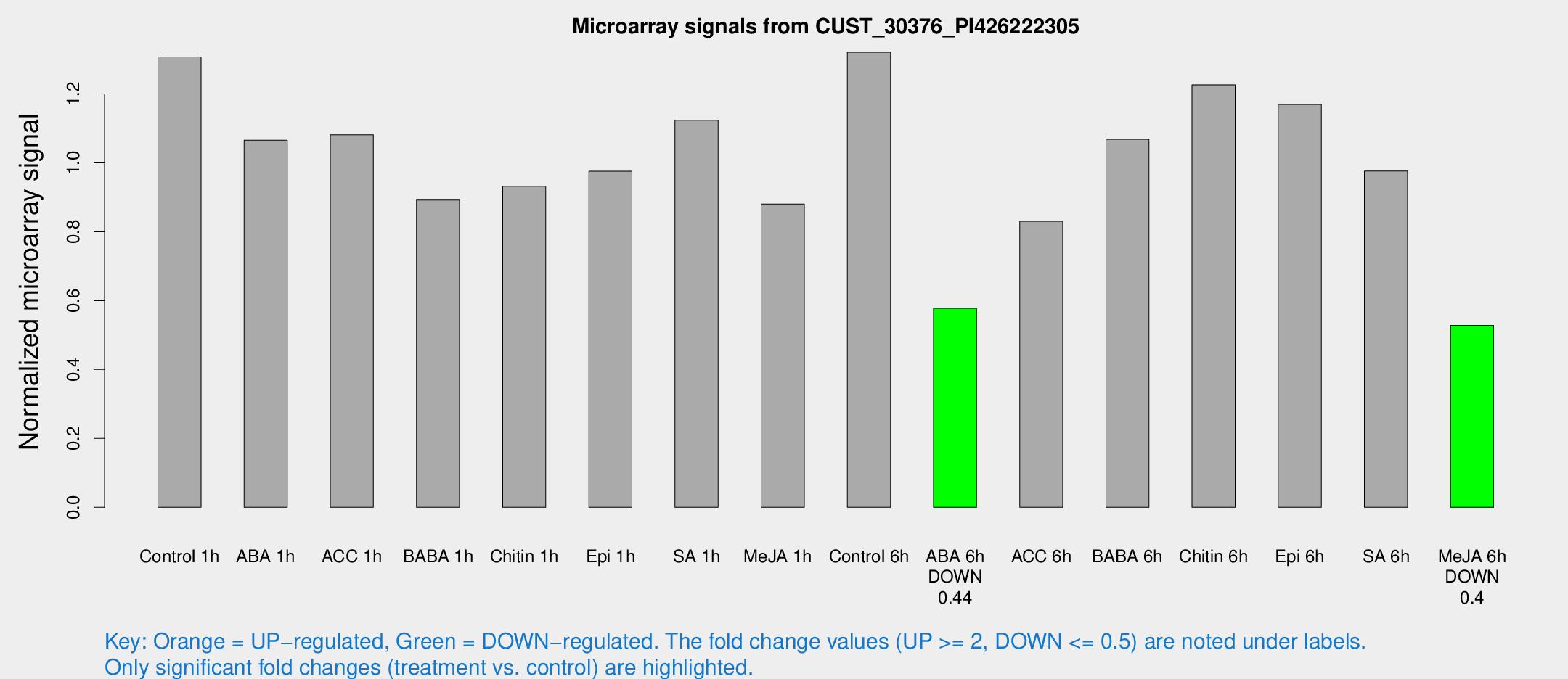
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 18645.3 | 2750.7 | 1.30713 | 0.103103 |
| ABA 1h | 13308.3 | 1351.87 | 1.06581 | 0.103073 |
| ACC 1h | 15718.1 | 1796.45 | 1.08125 | 0.0624268 |
| BABA 1h | 13004.7 | 3302.72 | 0.891872 | 0.17765 |
| Chitin 1h | 11755.3 | 766.224 | 0.931995 | 0.0538093 |
| Epi 1h | 11830 | 685.158 | 0.975448 | 0.0563181 |
| SA 1h | 16264.2 | 1493.77 | 1.12349 | 0.0648652 |
| Me-JA 1h | 10131.5 | 929.835 | 0.880547 | 0.0508391 |
| Control 6h | 19675.2 | 4546.66 | 1.32098 | 0.237119 |
| ABA 6h | 8624.52 | 607.274 | 0.577867 | 0.0333639 |
| ACC 6h | 13930.2 | 2904.98 | 0.830487 | 0.0921022 |
| BABA 6h | 17540.2 | 4004.87 | 1.06845 | 0.246954 |
| Chitin 6h | 18518.8 | 2384.43 | 1.22638 | 0.120447 |
| Epi 6h | 18594.1 | 1877.13 | 1.16941 | 0.136179 |
| SA 6h | 15288.9 | 4736.75 | 0.976259 | 0.26823 |
| Me-JA 6h | 8011.36 | 2157.88 | 0.528283 | 0.121748 |
Source Transcript PGSC0003DMT400069751 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G55800.1 | +1 | 2e-160 | 458 | 245/310 (79%) | sedoheptulose-bisphosphatase | chr3:20709640-20711421 FORWARD LENGTH=393 |
