Probe CUST_30258_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_30258_PI426222305 | JHI_St_60k_v1 | DMT400012160 | TGGTTGTTTTGATGAGAAACTCTTGCGCCAGAGAGAAAGCCCAAAATTTTCTAGTGTAAT |
All Microarray Probes Designed to Gene DMG400004766
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_30228_PI426222305 | JHI_St_60k_v1 | DMT400012159 | AAACTCTTGCGTGAAAGCTGGACCATCAGTTAGAGGAGCGGCATGGGGAAAACTTAATTT |
| CUST_30258_PI426222305 | JHI_St_60k_v1 | DMT400012160 | TGGTTGTTTTGATGAGAAACTCTTGCGCCAGAGAGAAAGCCCAAAATTTTCTAGTGTAAT |
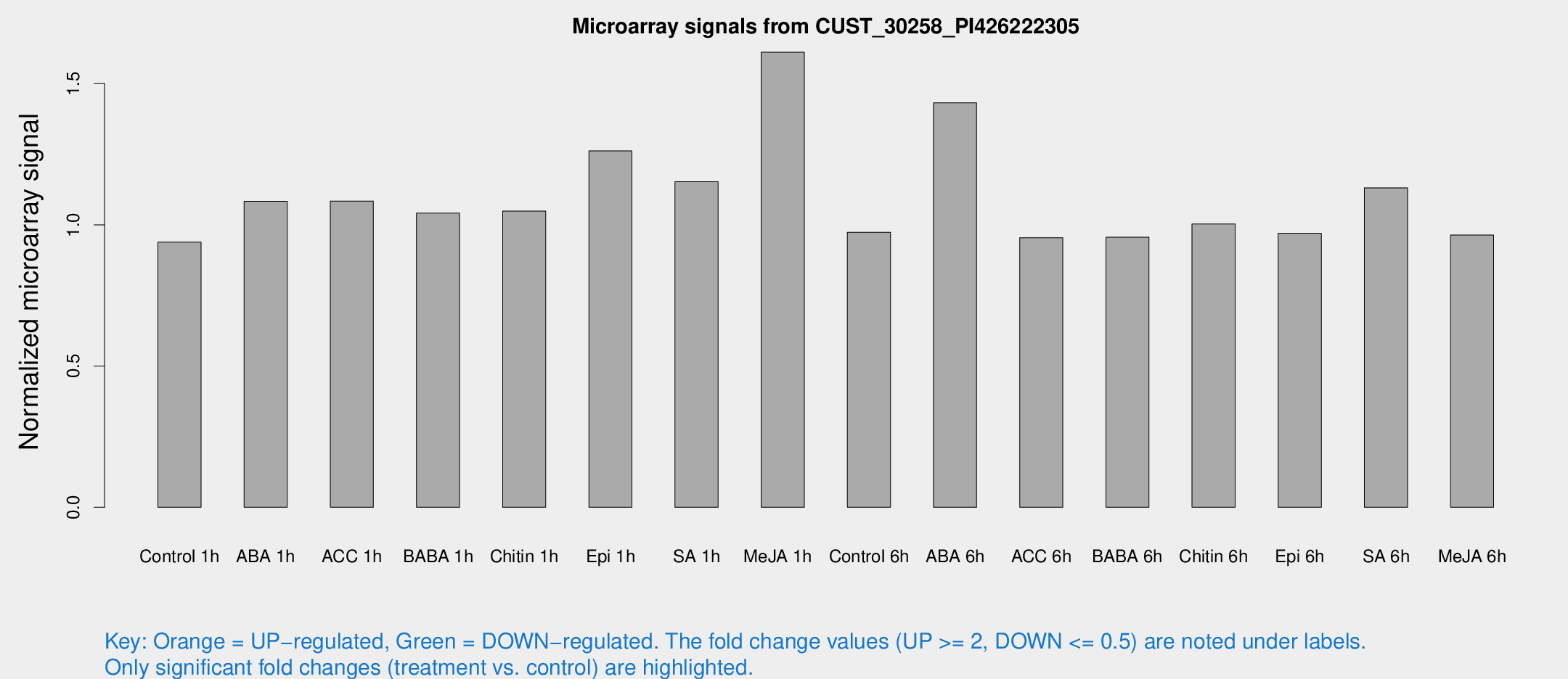
Microarray Signals from CUST_30258_PI426222305
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5.69974 | 3.30929 | 0.938805 | 0.54374 |
| ABA 1h | 5.82529 | 3.16231 | 1.08343 | 0.592244 |
| ACC 1h | 6.85288 | 4.04248 | 1.08415 | 0.628876 |
| BABA 1h | 6.09694 | 3.54172 | 1.04136 | 0.603115 |
| Chitin 1h | 5.69636 | 3.30359 | 1.04849 | 0.597884 |
| Epi 1h | 6.91971 | 3.22576 | 1.26161 | 0.631676 |
| SA 1h | 7.75122 | 3.31777 | 1.15248 | 0.563283 |
| Me-JA 1h | 8.1608 | 3.39927 | 1.61112 | 0.713725 |
| Control 6h | 5.91862 | 3.43346 | 0.973465 | 0.5638 |
| ABA 6h | 11.4706 | 5.58631 | 1.43213 | 0.661984 |
| ACC 6h | 6.7935 | 4.03902 | 0.954461 | 0.553464 |
| BABA 6h | 6.49662 | 3.77658 | 0.956347 | 0.554652 |
| Chitin 6h | 6.47315 | 3.75764 | 1.00298 | 0.581641 |
| Epi 6h | 6.71471 | 3.94522 | 0.970137 | 0.561979 |
| SA 6h | 6.78905 | 3.61811 | 1.13113 | 0.605507 |
| Me-JA 6h | 5.81372 | 3.38013 | 0.963765 | 0.560183 |
Source Transcript PGSC0003DMT400012160 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G13200.1 | +2 | 2e-76 | 248 | 112/174 (64%) | GRAM domain family protein | chr5:4207081-4208079 FORWARD LENGTH=272 |
