Probe CUST_29833_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_29833_PI426222305 | JHI_St_60k_v1 | DMT400033408 | ATTCCTTACAAGATCAGCCCCTCCACTGACTGCTCCAAGTACGTTATTTATATTTTGTGA |
All Microarray Probes Designed to Gene DMG400012837
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_29833_PI426222305 | JHI_St_60k_v1 | DMT400033408 | ATTCCTTACAAGATCAGCCCCTCCACTGACTGCTCCAAGTACGTTATTTATATTTTGTGA |
| CUST_29827_PI426222305 | JHI_St_60k_v1 | DMT400033409 | GCCGCTGGTATACCTAGAGTTTGTGGTCTAGTGCTATTAAAGGCATTAATGTGGGCAAAG |
Microarray Signals from CUST_29833_PI426222305
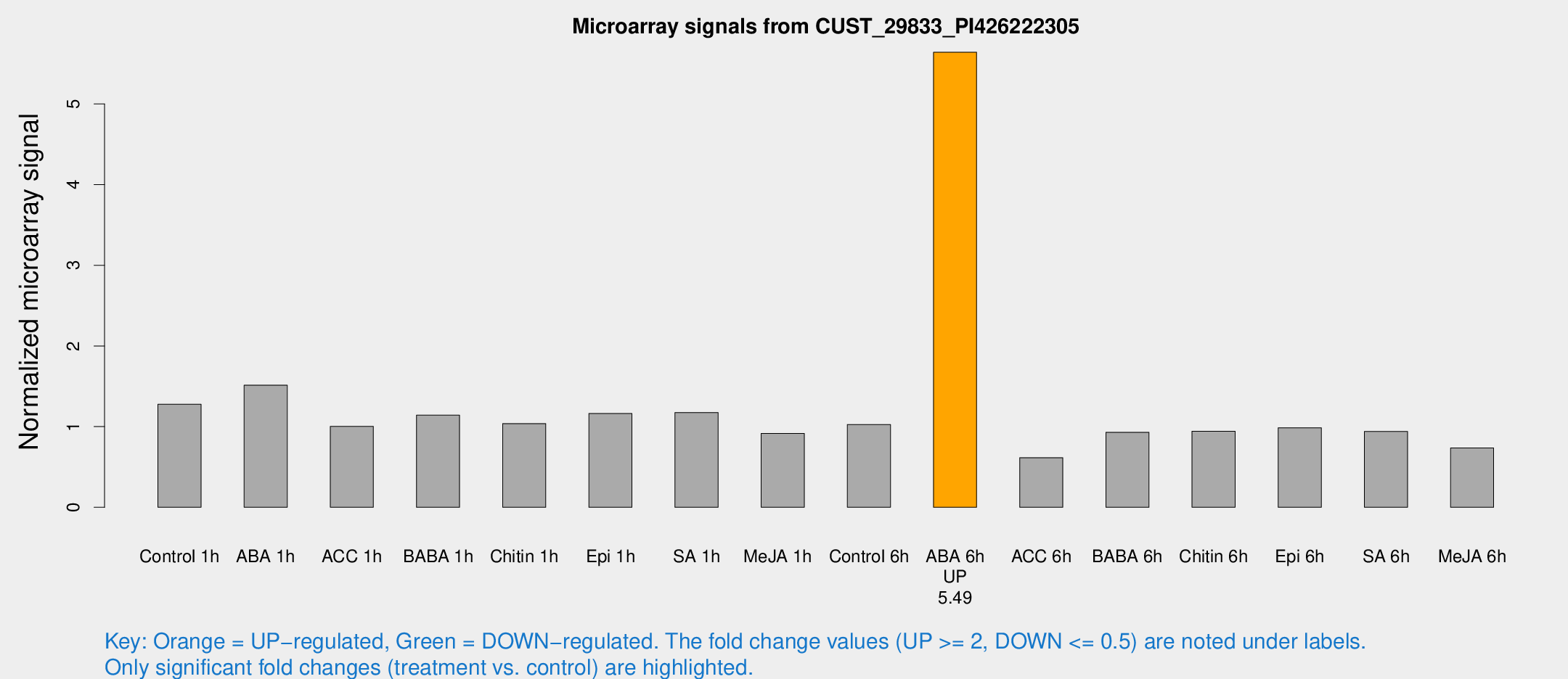
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 38494.6 | 2225.14 | 1.27795 | 0.0737826 |
| ABA 1h | 40659.2 | 3857.21 | 1.51376 | 0.12807 |
| ACC 1h | 35459.5 | 11515.8 | 1.00281 | 0.336232 |
| BABA 1h | 33348.3 | 2853.05 | 1.14194 | 0.222059 |
| Chitin 1h | 28228 | 2076.35 | 1.03751 | 0.126959 |
| Epi 1h | 31303.7 | 5452.45 | 1.1629 | 0.212463 |
| SA 1h | 37398 | 5939.24 | 1.17531 | 0.282773 |
| Me-JA 1h | 22722.1 | 2215.78 | 0.91646 | 0.153116 |
| Control 6h | 33349.3 | 8132.27 | 1.02672 | 0.201026 |
| ABA 6h | 181765 | 15060.5 | 5.64104 | 0.325686 |
| ACC 6h | 21928.5 | 3672.95 | 0.615328 | 0.131654 |
| BABA 6h | 31460.4 | 1976.49 | 0.930282 | 0.05371 |
| Chitin 6h | 30381.9 | 2328.8 | 0.942961 | 0.0618822 |
| Epi 6h | 33603 | 2326.85 | 0.98599 | 0.142273 |
| SA 6h | 28132 | 2199.59 | 0.939464 | 0.056611 |
| Me-JA 6h | 23891.4 | 7155.55 | 0.735489 | 0.161516 |
Source Transcript PGSC0003DMT400033408 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G38540.1 | +1 | 3e-23 | 89 | 58/111 (52%) | lipid transfer protein 1 | chr2:16130418-16130893 FORWARD LENGTH=118 |
