Probe CUST_29418_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_29418_PI426222305 | JHI_St_60k_v1 | DMT400075869 | GGACCACTTAAGATTTCTAGCTCCCAATAAAGTTTGTAACATTCAAATGCTCATCTATCA |
All Microarray Probes Designed to Gene DMG400029498
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_29418_PI426222305 | JHI_St_60k_v1 | DMT400075869 | GGACCACTTAAGATTTCTAGCTCCCAATAAAGTTTGTAACATTCAAATGCTCATCTATCA |
| CUST_29477_PI426222305 | JHI_St_60k_v1 | DMT400075870 | ATTCCGGAAATATTATCATGTAAAATTCTGCTATCACACTTTGGGAGGACTTCAAGATCA |
Microarray Signals from CUST_29418_PI426222305
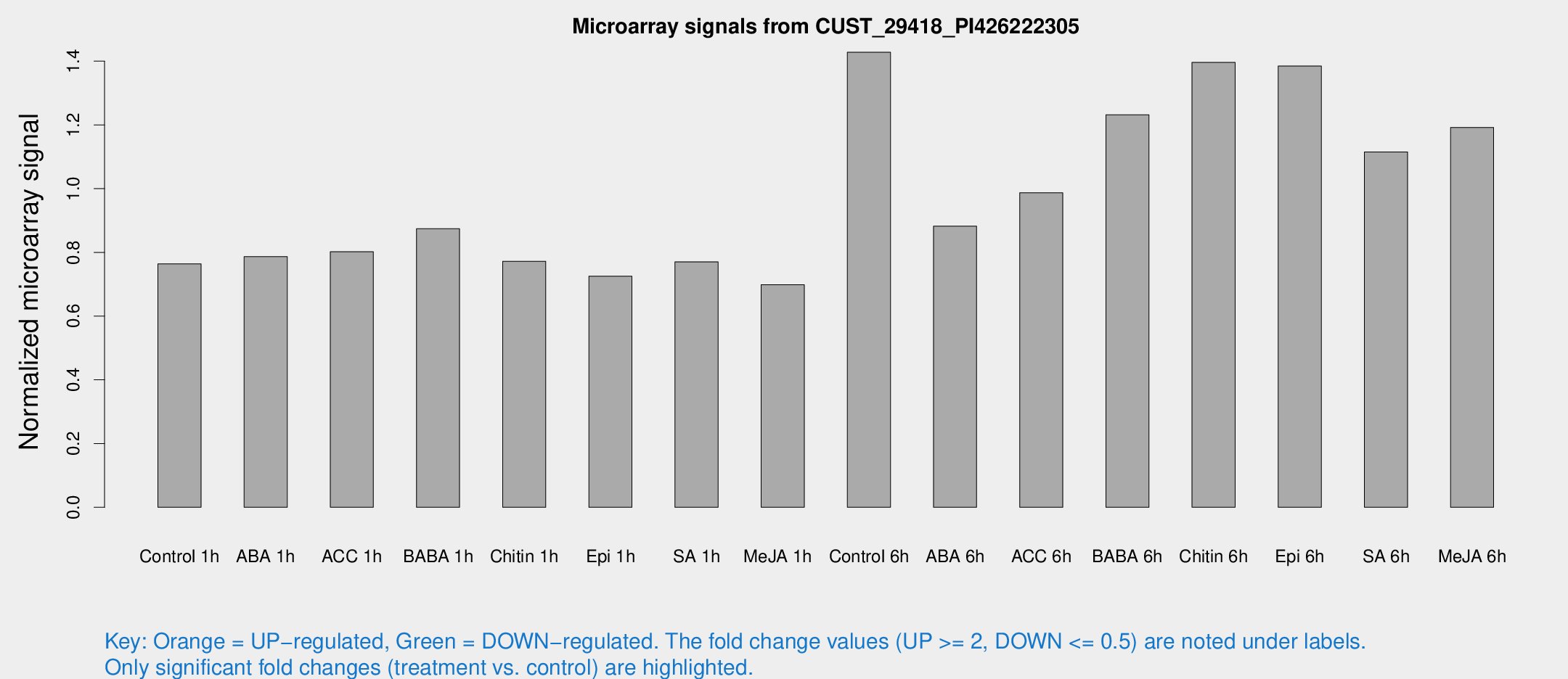
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 67.1174 | 14.2089 | 0.764095 | 0.140204 |
| ABA 1h | 59.3996 | 7.5838 | 0.786642 | 0.131875 |
| ACC 1h | 73.4367 | 18.3262 | 0.801791 | 0.161634 |
| BABA 1h | 74.0357 | 14.3846 | 0.874126 | 0.102237 |
| Chitin 1h | 60.8211 | 13.0129 | 0.772012 | 0.104327 |
| Epi 1h | 58.4978 | 19.4836 | 0.725239 | 0.222979 |
| SA 1h | 67.4293 | 8.18522 | 0.770258 | 0.0668042 |
| Me-JA 1h | 51.6521 | 14.8714 | 0.698692 | 0.13869 |
| Control 6h | 129.766 | 33.6809 | 1.42801 | 0.280439 |
| ABA 6h | 86.909 | 23.8036 | 0.882287 | 0.331837 |
| ACC 6h | 99.1715 | 18.5018 | 0.987039 | 0.157023 |
| BABA 6h | 118.769 | 18.381 | 1.23171 | 0.165667 |
| Chitin 6h | 125.644 | 9.97068 | 1.39615 | 0.0885504 |
| Epi 6h | 131.736 | 8.76593 | 1.38449 | 0.190372 |
| SA 6h | 96.6611 | 18.8379 | 1.11479 | 0.118617 |
| Me-JA 6h | 107.319 | 29.6238 | 1.19203 | 0.228638 |
Source Transcript PGSC0003DMT400075869 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G01910.1 | +3 | 0.0 | 734 | 396/591 (67%) | Microtubule associated protein (MAP65/ASE1) family protein | chr2:417191-420182 FORWARD LENGTH=608 |
