Probe CUST_28982_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28982_PI426222305 | JHI_St_60k_v1 | DMT400030514 | CGAGAGAACGAATTAAGGCAACGTGAATTGTCAATCTTTCATAGAAATTTCATTTGTGCT |
All Microarray Probes Designed to Gene DMG400011686
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28982_PI426222305 | JHI_St_60k_v1 | DMT400030514 | CGAGAGAACGAATTAAGGCAACGTGAATTGTCAATCTTTCATAGAAATTTCATTTGTGCT |
| CUST_29086_PI426222305 | JHI_St_60k_v1 | DMT400030515 | AAGCAGGTCGTGGATAATGTTTACAAGGCGGCTTCAGAGTATGTTGATTTTCAAAACAAG |
Microarray Signals from CUST_28982_PI426222305
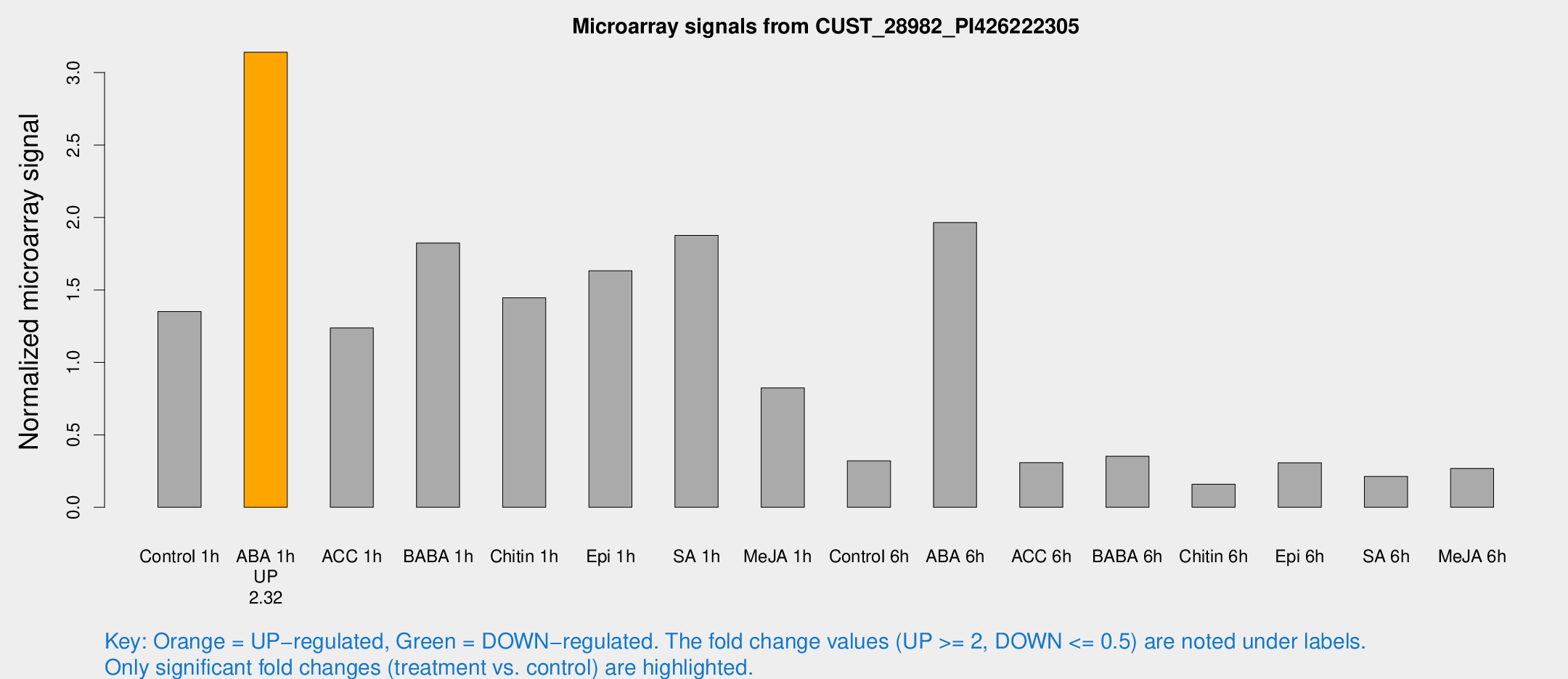
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 101.851 | 9.03664 | 1.35127 | 0.179423 |
| ABA 1h | 208.182 | 12.4445 | 3.14018 | 0.375844 |
| ACC 1h | 98.5883 | 18.2923 | 1.23744 | 0.150358 |
| BABA 1h | 133.464 | 17.0574 | 1.82344 | 0.245382 |
| Chitin 1h | 101.337 | 21.6491 | 1.44537 | 0.336913 |
| Epi 1h | 106.071 | 6.90682 | 1.63281 | 0.144812 |
| SA 1h | 154.501 | 36.9004 | 1.87576 | 0.454425 |
| Me-JA 1h | 50.329 | 4.43509 | 0.824151 | 0.0724862 |
| Control 6h | 44.2871 | 30.9259 | 0.321277 | 0.369842 |
| ABA 6h | 171.616 | 52.1283 | 1.96526 | 0.519857 |
| ACC 6h | 30.9639 | 11.2234 | 0.30778 | 0.15858 |
| BABA 6h | 31.4214 | 7.11654 | 0.352489 | 0.104932 |
| Chitin 6h | 15.5177 | 6.98915 | 0.159261 | 0.0809794 |
| Epi 6h | 39.0962 | 22.7703 | 0.306899 | 0.332155 |
| SA 6h | 19.0163 | 7.06358 | 0.212942 | 0.0828243 |
| Me-JA 6h | 20.1841 | 3.5388 | 0.268449 | 0.048452 |
Source Transcript PGSC0003DMT400030514 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G47710.1 | +2 | 7e-96 | 303 | 155/261 (59%) | Serine protease inhibitor (SERPIN) family protein | chr1:17558271-17560061 FORWARD LENGTH=391 |
