Probe CUST_28781_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28781_PI426222305 | JHI_St_60k_v1 | DMT400010401 | TCTAATCTGGTTTCACCATGGATTATGAATAAAGATTGGCAGATGAACTATCATGTCCAT |
All Microarray Probes Designed to Gene DMG400004064
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28844_PI426222305 | JHI_St_60k_v1 | DMT400010400 | CGGGTTCGTCTCTTTGTAGATGAATCTGGAAAAGTTGCTCAAGAGCCTAGACTTGGCTAA |
| CUST_28781_PI426222305 | JHI_St_60k_v1 | DMT400010401 | TCTAATCTGGTTTCACCATGGATTATGAATAAAGATTGGCAGATGAACTATCATGTCCAT |
Microarray Signals from CUST_28781_PI426222305
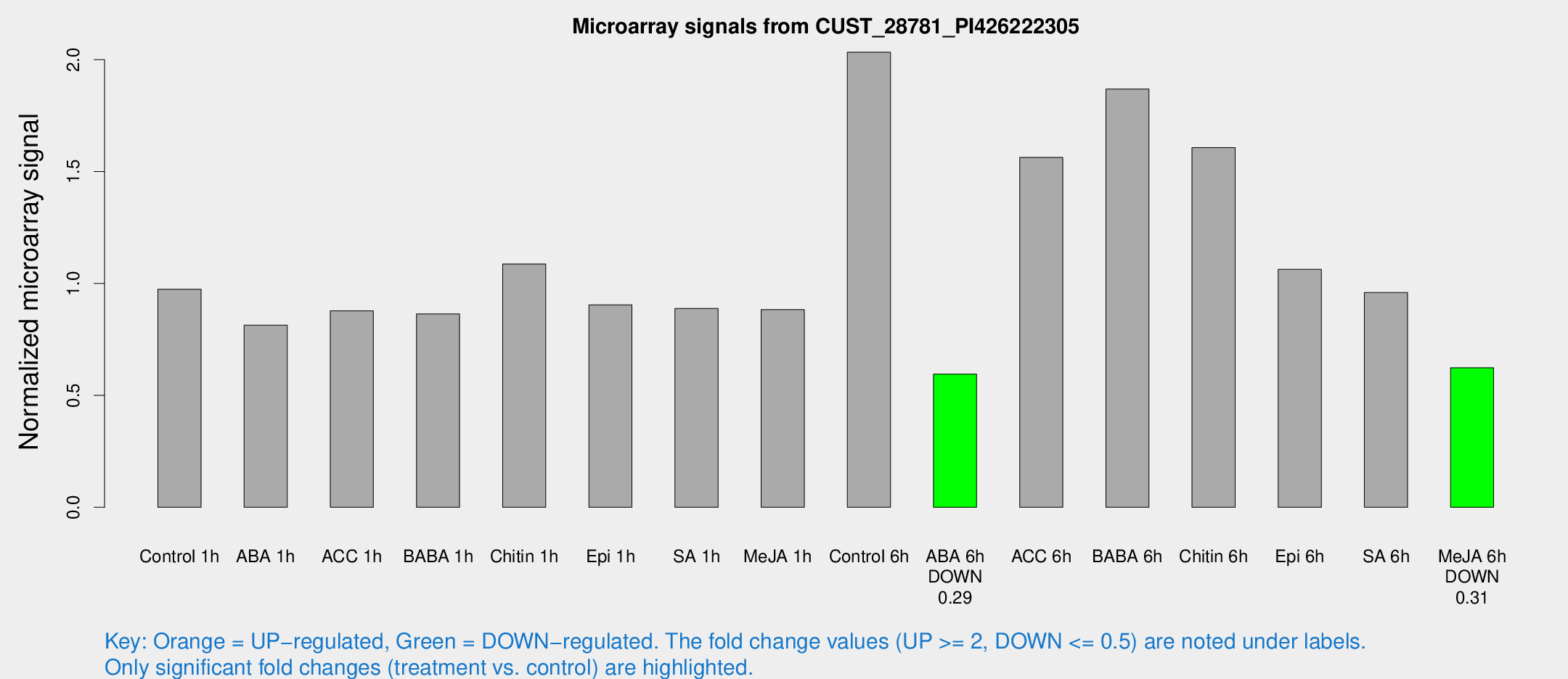
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 527.775 | 62.0307 | 0.974538 | 0.0810157 |
| ABA 1h | 387.381 | 31.0373 | 0.813856 | 0.0897739 |
| ACC 1h | 503.207 | 96.2963 | 0.877731 | 0.125171 |
| BABA 1h | 445.745 | 25.9784 | 0.863895 | 0.128556 |
| Chitin 1h | 542.106 | 108.346 | 1.08725 | 0.216129 |
| Epi 1h | 419.553 | 24.4607 | 0.905013 | 0.052659 |
| SA 1h | 493.545 | 50.9088 | 0.888753 | 0.146483 |
| Me-JA 1h | 386.413 | 22.59 | 0.883322 | 0.0989296 |
| Control 6h | 1221.24 | 369.496 | 2.03346 | 0.560172 |
| ABA 6h | 359.984 | 87.869 | 0.595166 | 0.177759 |
| ACC 6h | 996.665 | 206.345 | 1.56311 | 0.0904482 |
| BABA 6h | 1132.21 | 124.269 | 1.86862 | 0.262967 |
| Chitin 6h | 932.344 | 134.709 | 1.60735 | 0.23236 |
| Epi 6h | 662.558 | 112.329 | 1.06317 | 0.285841 |
| SA 6h | 531.902 | 118.839 | 0.959478 | 0.147958 |
| Me-JA 6h | 335.34 | 33.0123 | 0.623539 | 0.0364887 |
Source Transcript PGSC0003DMT400010401 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | None | - | - | - | - | - |
