Probe CUST_28489_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28489_PI426222305 | JHI_St_60k_v1 | DMT400009949 | CTGGAGGAAAACTATTGAATTCTGTTGAACTCGTATCTTGTACTGTAGAATACAAGAAGA |
All Microarray Probes Designed to Gene DMG400003897
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28489_PI426222305 | JHI_St_60k_v1 | DMT400009949 | CTGGAGGAAAACTATTGAATTCTGTTGAACTCGTATCTTGTACTGTAGAATACAAGAAGA |
| CUST_28488_PI426222305 | JHI_St_60k_v1 | DMT400009950 | CTGGAGGAAAACTATTGAATTCTGTTGAACTCGTATCTTGTACTGTAGAATACAAGAAGA |
Microarray Signals from CUST_28489_PI426222305
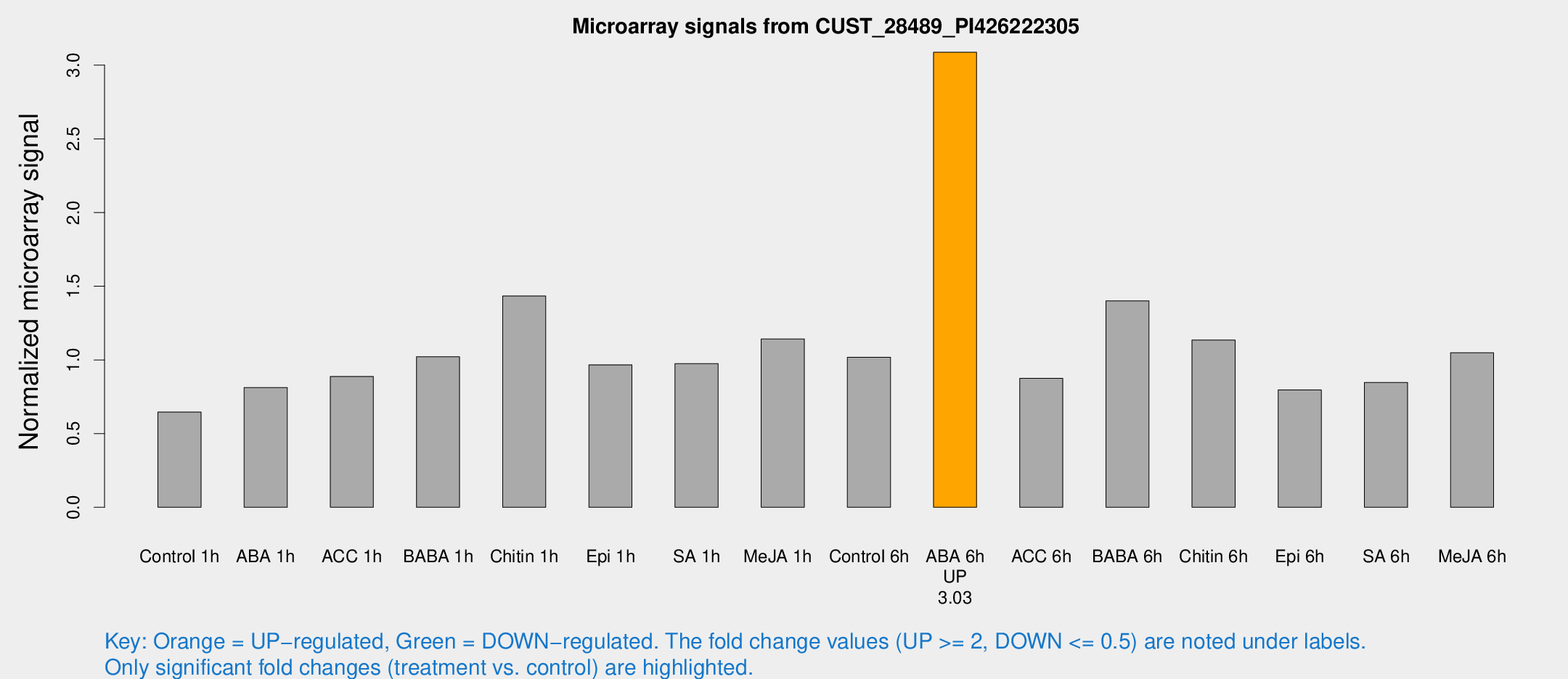
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 137.803 | 17.793 | 0.645979 | 0.0409196 |
| ABA 1h | 158.31 | 31.9241 | 0.812342 | 0.139922 |
| ACC 1h | 194.452 | 24.5142 | 0.887892 | 0.0560775 |
| BABA 1h | 220.994 | 52.3574 | 1.02119 | 0.175893 |
| Chitin 1h | 277.235 | 41.8361 | 1.4336 | 0.178624 |
| Epi 1h | 177.258 | 17.4308 | 0.966944 | 0.0665706 |
| SA 1h | 211.44 | 16.5115 | 0.974737 | 0.0585606 |
| Me-JA 1h | 198.664 | 23.2558 | 1.14212 | 0.0775026 |
| Control 6h | 236.171 | 63.598 | 1.01778 | 0.25473 |
| ABA 6h | 700.047 | 84.7862 | 3.08773 | 0.400671 |
| ACC 6h | 220.245 | 46.9258 | 0.875177 | 0.0729264 |
| BABA 6h | 331.926 | 31.0078 | 1.40112 | 0.0826652 |
| Chitin 6h | 254.636 | 16.3328 | 1.1353 | 0.0823699 |
| Epi 6h | 191.521 | 25.267 | 0.795971 | 0.130026 |
| SA 6h | 178.211 | 20.2075 | 0.84721 | 0.0523324 |
| Me-JA 6h | 238.826 | 60.7369 | 1.04923 | 0.254712 |
Source Transcript PGSC0003DMT400009949 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G22920.1 | +3 | 1e-151 | 438 | 183/281 (65%) | CHY-type/CTCHY-type/RING-type Zinc finger protein | chr5:7665143-7667031 FORWARD LENGTH=291 |
