Probe CUST_28483_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28483_PI426222305 | JHI_St_60k_v1 | DMT400009959 | CCAGATAGCATTTGAGCTTCGAAGACCTCATGGCCTTCTAGATATATTTACTGAACTCCA |
All Microarray Probes Designed to Gene DMG401003900
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28483_PI426222305 | JHI_St_60k_v1 | DMT400009959 | CCAGATAGCATTTGAGCTTCGAAGACCTCATGGCCTTCTAGATATATTTACTGAACTCCA |
| CUST_28537_PI426222305 | JHI_St_60k_v1 | DMT400009957 | TCCGCCAATTGAAATTGTAAGACAAAGCAATATGCACAATTTGTTAGACGACAAGGGTAT |
Microarray Signals from CUST_28483_PI426222305
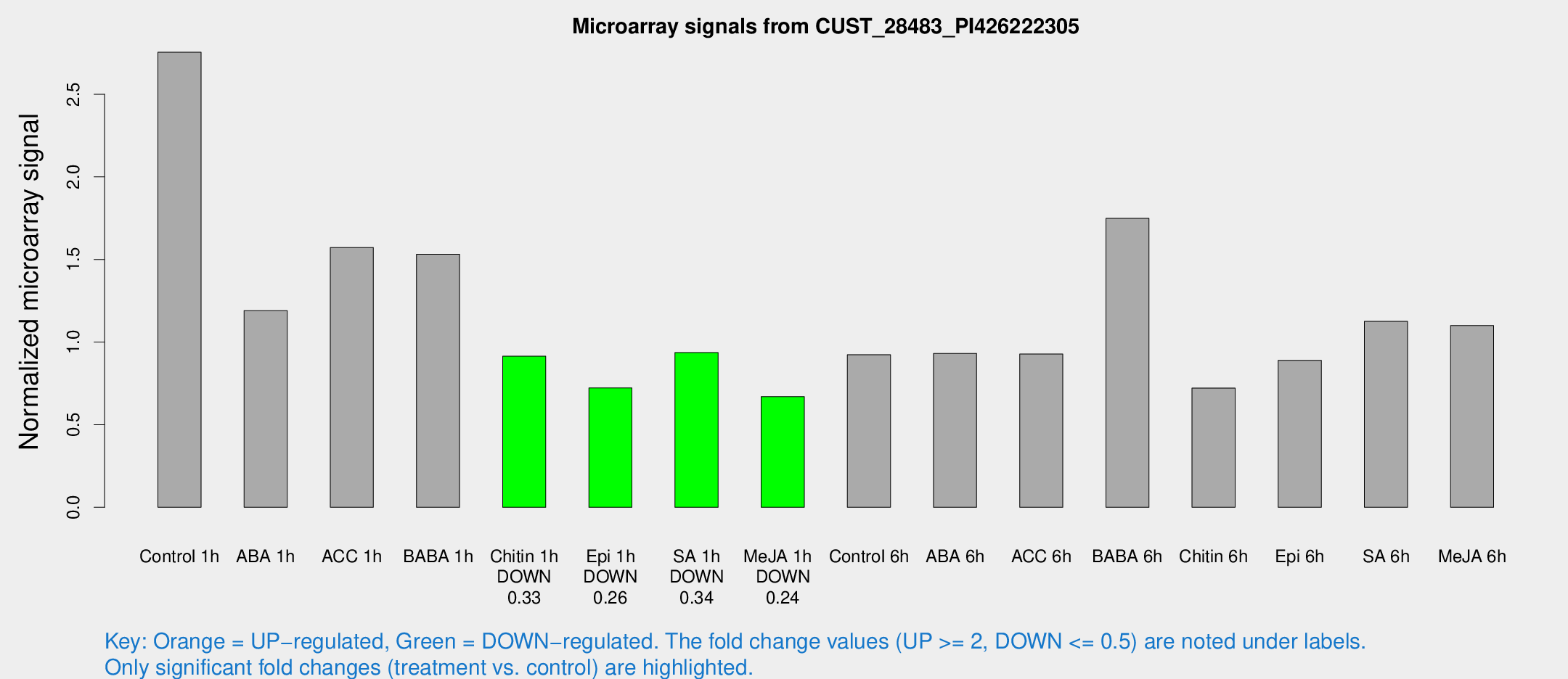
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 27.3919 | 3.40627 | 2.75441 | 0.347429 |
| ABA 1h | 10.9656 | 3.02967 | 1.19071 | 0.371221 |
| ACC 1h | 16.3117 | 3.44257 | 1.57231 | 0.34779 |
| BABA 1h | 14.6579 | 3.32703 | 1.53188 | 0.351754 |
| Chitin 1h | 8.70796 | 3.14117 | 0.914708 | 0.36846 |
| Epi 1h | 6.33652 | 3.05538 | 0.722296 | 0.360051 |
| SA 1h | 9.90112 | 3.10874 | 0.936494 | 0.321941 |
| Me-JA 1h | 5.4311 | 3.15822 | 0.669837 | 0.388113 |
| Control 6h | 9.757 | 3.29363 | 0.923621 | 0.363267 |
| ABA 6h | 10.0951 | 3.38583 | 0.931278 | 0.331234 |
| ACC 6h | 11.5968 | 3.94529 | 0.927784 | 0.366747 |
| BABA 6h | 20.7616 | 5.06584 | 1.74872 | 0.440705 |
| Chitin 6h | 7.71101 | 3.56866 | 0.721911 | 0.344637 |
| Epi 6h | 10.7595 | 3.7353 | 0.889769 | 0.368019 |
| SA 6h | 11.5189 | 3.46526 | 1.12582 | 0.3917 |
| Me-JA 6h | 12.0899 | 3.38691 | 1.09997 | 0.393679 |
Source Transcript PGSC0003DMT400009959 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G16750.1 | +3 | 0.0 | 247 | 116/173 (67%) | Transducin family protein / WD-40 repeat family protein | chr5:5504541-5509266 REVERSE LENGTH=876 |
