Probe CUST_28475_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28475_PI426222305 | JHI_St_60k_v1 | DMT400009955 | AATTGTTGCAGAAGAAGAGTCTCTGGAGGATGAAATATCACCCCAGGGCCTTGTTTTGGT |
All Microarray Probes Designed to Gene DMG401003899
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28505_PI426222305 | JHI_St_60k_v1 | DMT400009953 | CGACGACTTCAAGTGAATATACAATAATAACTGTGTATTAGATCATAGTAGACGACAGTC |
| CUST_28475_PI426222305 | JHI_St_60k_v1 | DMT400009955 | AATTGTTGCAGAAGAAGAGTCTCTGGAGGATGAAATATCACCCCAGGGCCTTGTTTTGGT |
Microarray Signals from CUST_28475_PI426222305
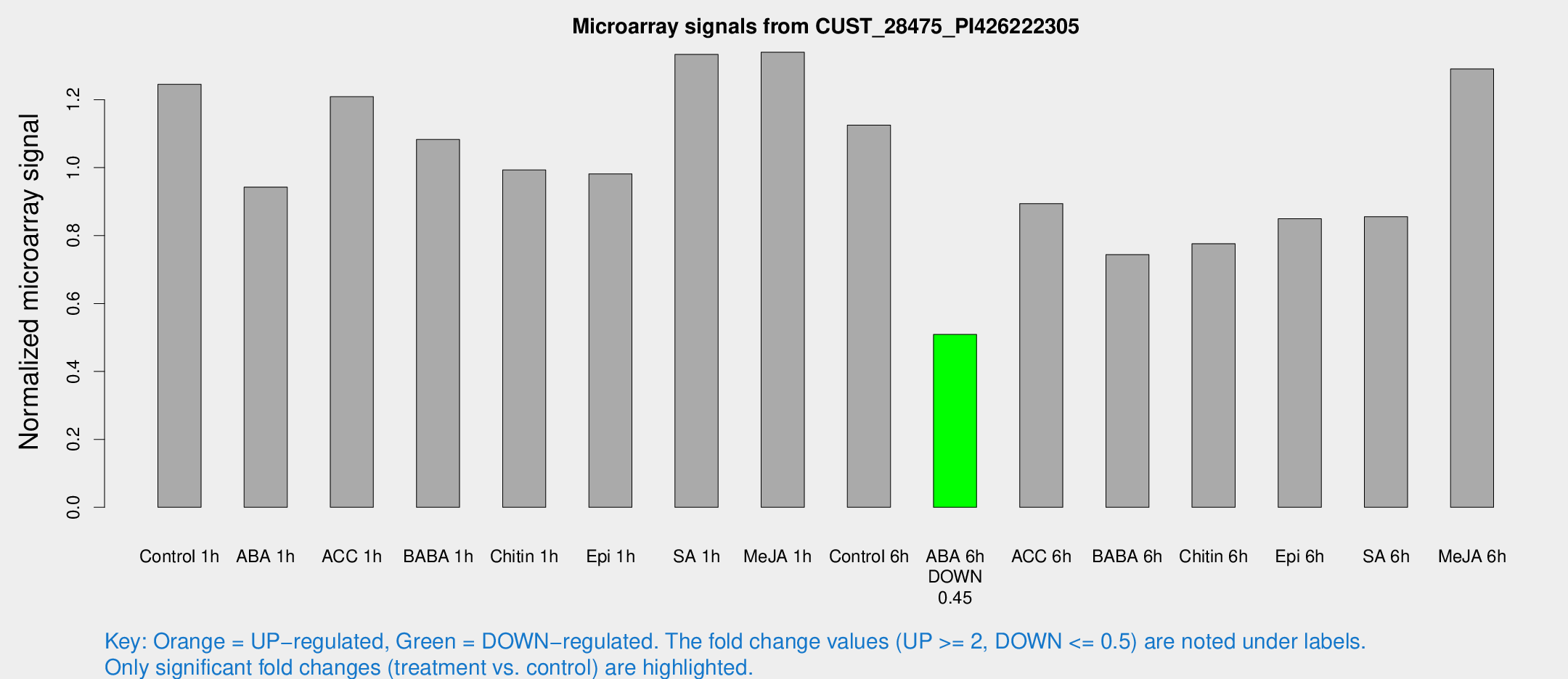
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 1537.26 | 277.18 | 1.24559 | 0.141923 |
| ABA 1h | 1008.31 | 100.866 | 0.942664 | 0.054495 |
| ACC 1h | 1507.07 | 168.481 | 1.209 | 0.0698735 |
| BABA 1h | 1304.83 | 251.238 | 1.08281 | 0.121875 |
| Chitin 1h | 1084.99 | 123.389 | 0.993184 | 0.0608655 |
| Epi 1h | 1024.31 | 79.176 | 0.981838 | 0.0699254 |
| SA 1h | 1663.12 | 198.14 | 1.33333 | 0.151917 |
| Me-JA 1h | 1328.38 | 151.298 | 1.33985 | 0.0774215 |
| Control 6h | 1428.3 | 304.511 | 1.12516 | 0.1784 |
| ABA 6h | 651.656 | 47.0483 | 0.508928 | 0.0294918 |
| ACC 6h | 1245.88 | 151.765 | 0.894156 | 0.0516885 |
| BABA 6h | 1000.87 | 59.9281 | 0.743879 | 0.0433327 |
| Chitin 6h | 990.807 | 57.3741 | 0.776171 | 0.0448933 |
| Epi 6h | 1174.09 | 182.955 | 0.849862 | 0.127991 |
| SA 6h | 1082.61 | 255.06 | 0.855744 | 0.132331 |
| Me-JA 6h | 1555.81 | 154.463 | 1.29069 | 0.0745646 |
Source Transcript PGSC0003DMT400009955 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G09920.1 | +1 | 3e-150 | 448 | 230/299 (77%) | phosphatidyl inositol monophosphate 5 kinase | chr3:3040426-3043676 REVERSE LENGTH=815 |
