Probe CUST_28318_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28318_PI426222305 | JHI_St_60k_v1 | DMT400044325 | GATGATAAAAGATGGAAGGTTCTCAAGATAAGCCATACTCAGCTACTTATCCTCTTTATA |
All Microarray Probes Designed to Gene DMG400017211
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28305_PI426222305 | JHI_St_60k_v1 | DMT400044324 | GATGATAAAAGATGGAAGGTTCTCAAGATAAGCCATACTCAGCTACTTATCCTCTTTATA |
| CUST_28318_PI426222305 | JHI_St_60k_v1 | DMT400044325 | GATGATAAAAGATGGAAGGTTCTCAAGATAAGCCATACTCAGCTACTTATCCTCTTTATA |
Microarray Signals from CUST_28318_PI426222305
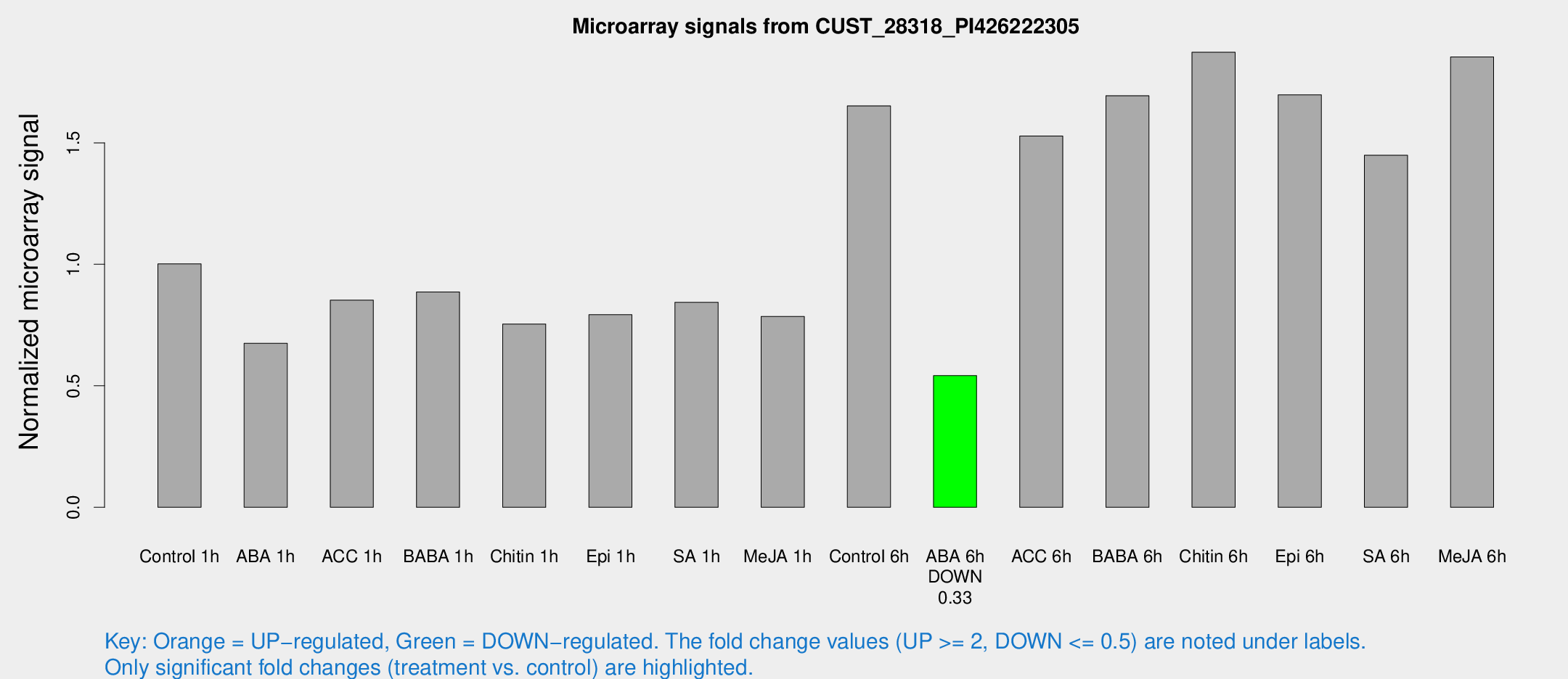
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 465.231 | 47.2402 | 1.00202 | 0.148497 |
| ABA 1h | 275.656 | 18.3971 | 0.675227 | 0.0833512 |
| ACC 1h | 424.577 | 88.5174 | 0.852795 | 0.14247 |
| BABA 1h | 397.836 | 45.4856 | 0.886144 | 0.0519265 |
| Chitin 1h | 312.497 | 18.4637 | 0.754319 | 0.0986074 |
| Epi 1h | 319.546 | 38.2913 | 0.792706 | 0.099774 |
| SA 1h | 400.514 | 30.4115 | 0.843466 | 0.0492996 |
| Me-JA 1h | 295.113 | 17.4711 | 0.78554 | 0.0464285 |
| Control 6h | 831.142 | 216.683 | 1.65197 | 0.385765 |
| ABA 6h | 267.297 | 26.893 | 0.541732 | 0.0323379 |
| ACC 6h | 833.688 | 160.921 | 1.52769 | 0.10795 |
| BABA 6h | 882.541 | 100.358 | 1.69382 | 0.158862 |
| Chitin 6h | 918.172 | 53.301 | 1.87282 | 0.108482 |
| Epi 6h | 895.679 | 117.146 | 1.69758 | 0.371456 |
| SA 6h | 660.206 | 38.4381 | 1.44867 | 0.254147 |
| Me-JA 6h | 858.034 | 88.589 | 1.8536 | 0.10737 |
Source Transcript PGSC0003DMT400044325 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G23410.1 | +1 | 0.0 | 762 | 427/726 (59%) | fatty alcohol oxidase 3 | chr3:8382860-8386024 FORWARD LENGTH=746 |
