Probe CUST_28250_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28250_PI426222305 | JHI_St_60k_v1 | DMT400044212 | GGTACAAGTCAAGGATTCTGGTACTAGCAGATATAATGTTGTTTCTGTTCATGTCTTTTT |
All Microarray Probes Designed to Gene DMG400017167
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28250_PI426222305 | JHI_St_60k_v1 | DMT400044212 | GGTACAAGTCAAGGATTCTGGTACTAGCAGATATAATGTTGTTTCTGTTCATGTCTTTTT |
Microarray Signals from CUST_28250_PI426222305
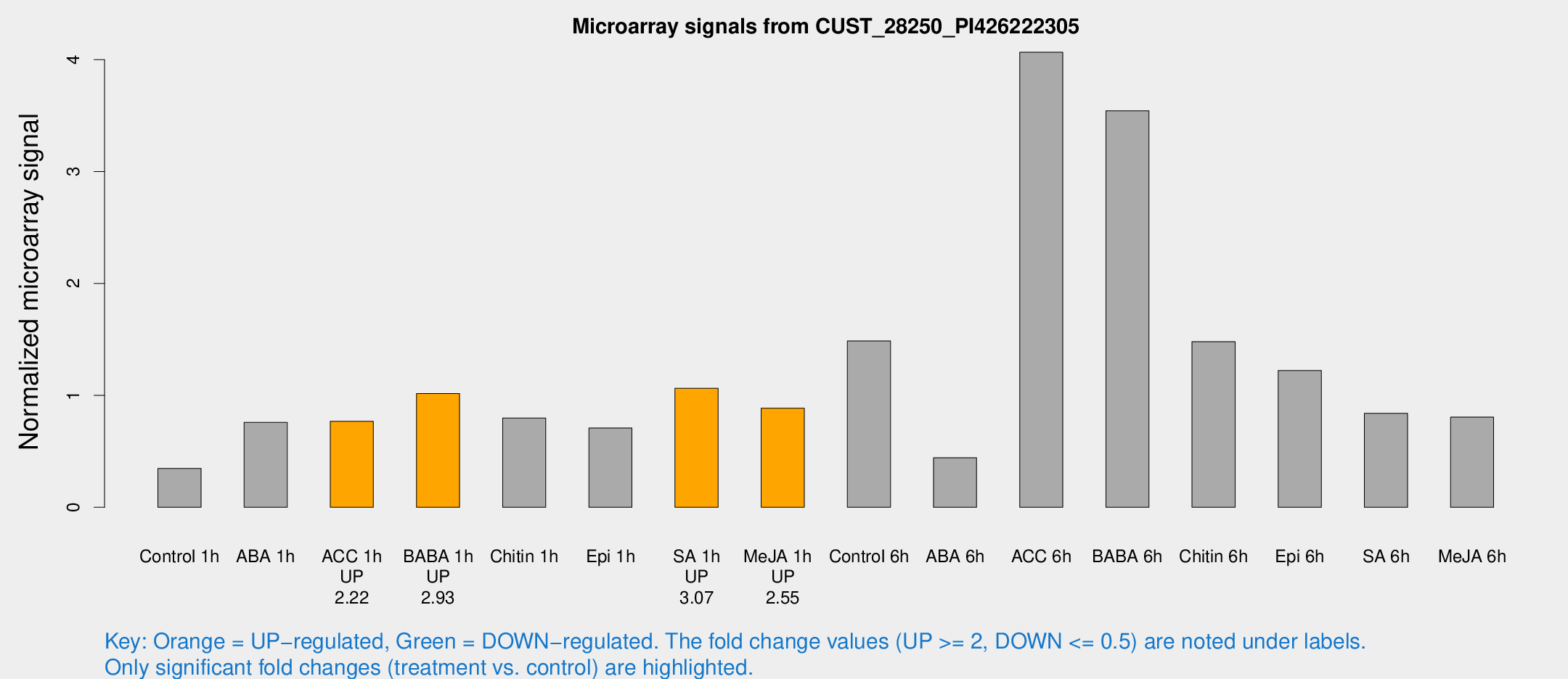
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 7.29003 | 3.7249 | 0.346789 | 0.177737 |
| ABA 1h | 16.4462 | 5.60517 | 0.758235 | 0.452129 |
| ACC 1h | 17.5822 | 4.63731 | 0.768593 | 0.261211 |
| BABA 1h | 21.1199 | 4.07821 | 1.01681 | 0.210269 |
| Chitin 1h | 17.7921 | 6.23553 | 0.797762 | 0.322106 |
| Epi 1h | 14.3292 | 4.11584 | 0.708313 | 0.296632 |
| SA 1h | 22.933 | 3.88748 | 1.06326 | 0.180028 |
| Me-JA 1h | 15.2078 | 3.90939 | 0.885664 | 0.227689 |
| Control 6h | 42.2052 | 16.7624 | 1.48645 | 1.00498 |
| ABA 6h | 11.188 | 4.16269 | 0.442485 | 0.207027 |
| ACC 6h | 107.832 | 31.9547 | 4.06649 | 0.8552 |
| BABA 6h | 101.406 | 47.0728 | 3.54365 | 1.60055 |
| Chitin 6h | 33.3276 | 4.59036 | 1.47958 | 0.206834 |
| Epi 6h | 31.9437 | 9.42816 | 1.22222 | 0.360072 |
| SA 6h | 17.8881 | 4.27813 | 0.839674 | 0.211127 |
| Me-JA 6h | 20.8222 | 8.25743 | 0.806285 | 0.37753 |
Source Transcript PGSC0003DMT400044212 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT2G41970.1 | +2 | 1e-134 | 399 | 201/309 (65%) | Protein kinase superfamily protein | chr2:17520517-17522304 REVERSE LENGTH=365 |
