Probe CUST_28180_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28180_PI426222305 | JHI_St_60k_v1 | DMT400004255 | GCATGTACTTTGAACTAGAGAGGATTCCTAGTCTAGCAGTGCATTATTTAGAATAGAAAT |
All Microarray Probes Designed to Gene DMG402001693
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28180_PI426222305 | JHI_St_60k_v1 | DMT400004255 | GCATGTACTTTGAACTAGAGAGGATTCCTAGTCTAGCAGTGCATTATTTAGAATAGAAAT |
Microarray Signals from CUST_28180_PI426222305
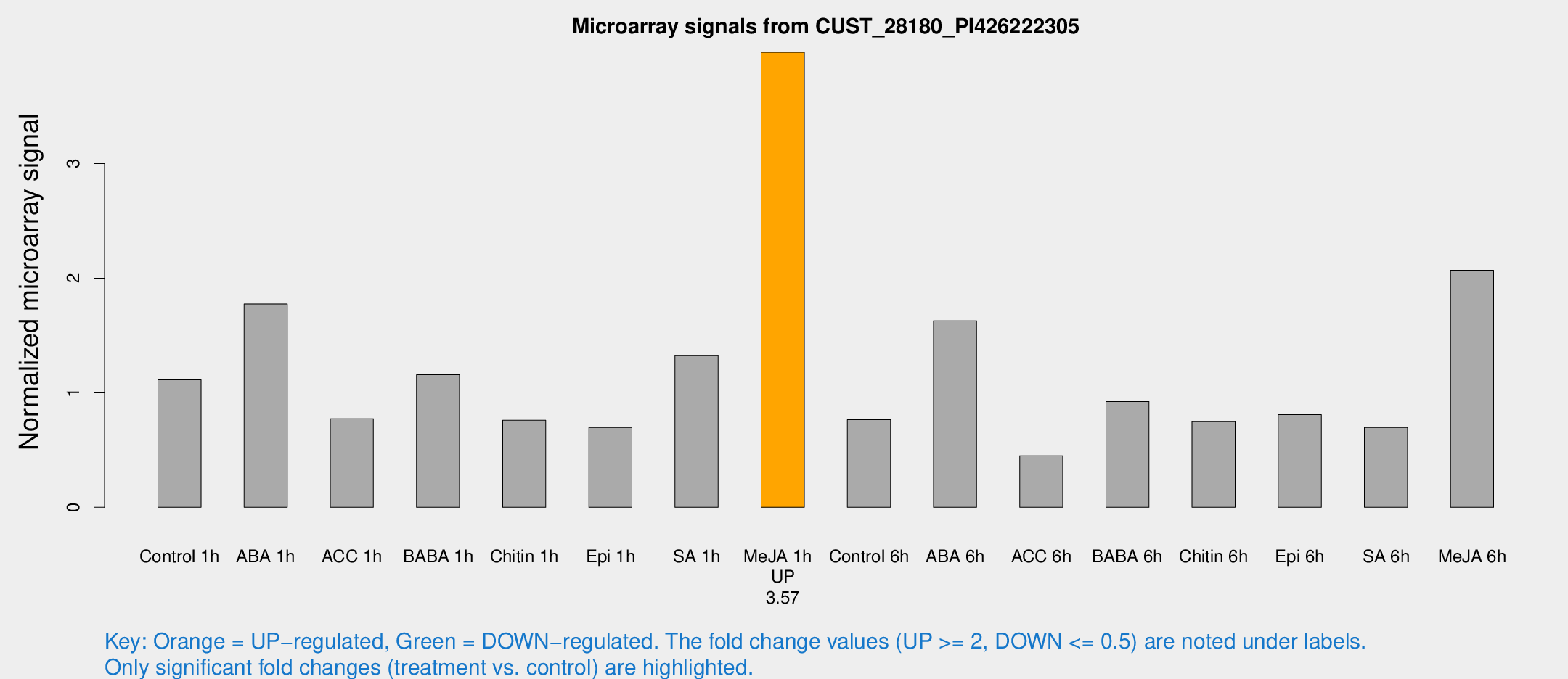
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 69.9373 | 5.26182 | 1.11252 | 0.133575 |
| ABA 1h | 103.765 | 24.6586 | 1.77506 | 0.285205 |
| ACC 1h | 56.5273 | 20.2255 | 0.773194 | 0.250815 |
| BABA 1h | 71.2557 | 9.5948 | 1.15756 | 0.210228 |
| Chitin 1h | 44.3372 | 8.28951 | 0.759582 | 0.122192 |
| Epi 1h | 40.0514 | 9.43439 | 0.697265 | 0.172848 |
| SA 1h | 100.438 | 41.375 | 1.32393 | 0.639037 |
| Me-JA 1h | 203.711 | 12.314 | 3.97179 | 0.239514 |
| Control 6h | 55.3599 | 18.6669 | 0.765347 | 0.24719 |
| ABA 6h | 108.627 | 7.33111 | 1.62822 | 0.109851 |
| ACC 6h | 33.3628 | 5.62628 | 0.450703 | 0.0650735 |
| BABA 6h | 70.2019 | 19.1483 | 0.924011 | 0.252671 |
| Chitin 6h | 52.365 | 12.1148 | 0.746859 | 0.166211 |
| Epi 6h | 57.6945 | 5.50145 | 0.809031 | 0.141241 |
| SA 6h | 43.7982 | 4.96614 | 0.697293 | 0.0757481 |
| Me-JA 6h | 130.441 | 12.7064 | 2.06866 | 0.261746 |
Source Transcript PGSC0003DMT400004255 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G28695.1 | +2 | 3e-108 | 323 | 169/290 (58%) | Nucleotide-diphospho-sugar transferase family protein | chr1:10081894-10083054 REVERSE LENGTH=329 |
