Probe CUST_28055_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28055_PI426222305 | JHI_St_60k_v1 | DMT400004192 | AGAGCTTGCTGATAATAGCAGTCGAAACATACTGCATACGAAAGAATTTAGGTAAGTTAT |
All Microarray Probes Designed to Gene DMG400001671
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_28112_PI426222305 | JHI_St_60k_v1 | DMT400004191 | CCAGCAGAACCTCAAGAAGAATGATATGTACCTAGCTATTCTTTGCTTTAGTAAAATTAC |
| CUST_28055_PI426222305 | JHI_St_60k_v1 | DMT400004192 | AGAGCTTGCTGATAATAGCAGTCGAAACATACTGCATACGAAAGAATTTAGGTAAGTTAT |
Microarray Signals from CUST_28055_PI426222305
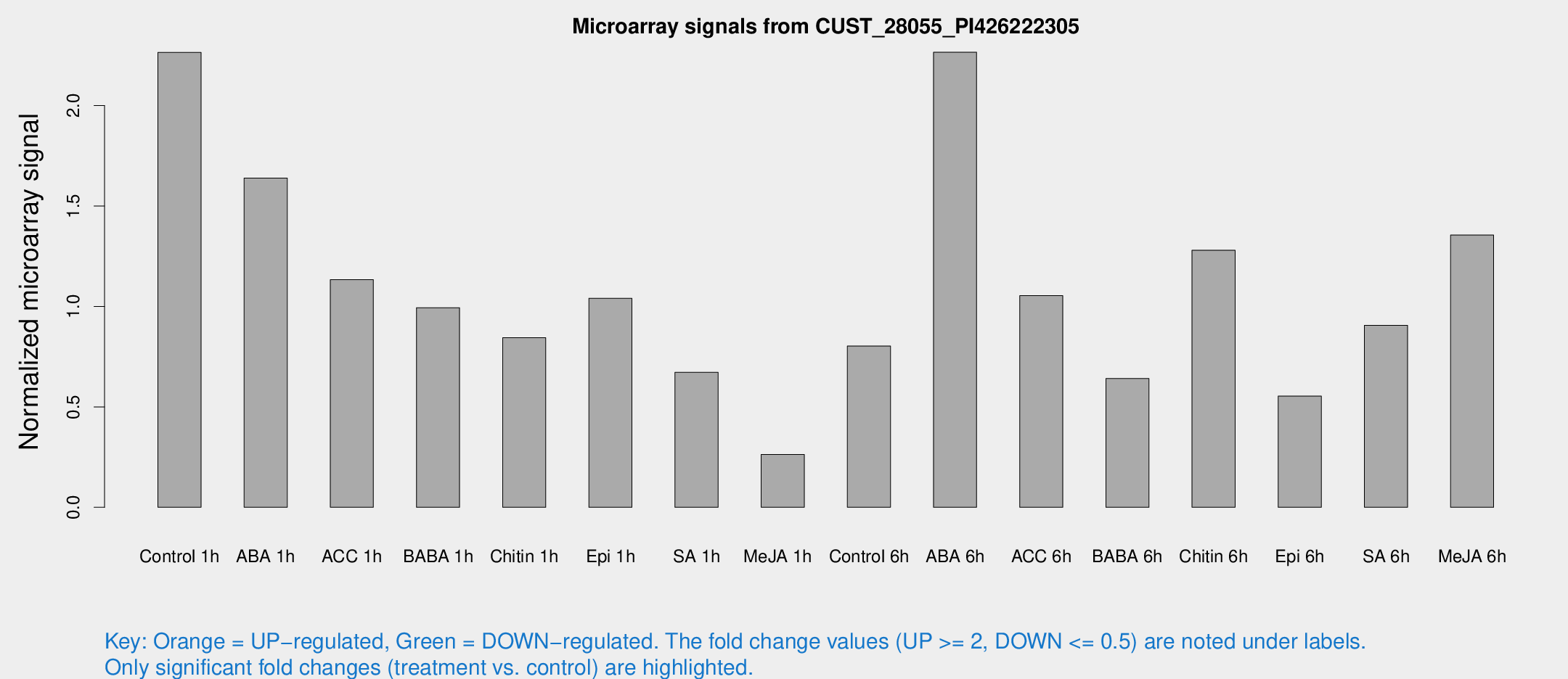
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 186.455 | 36.3297 | 2.26528 | 0.377223 |
| ABA 1h | 116.617 | 17.7275 | 1.63878 | 0.23973 |
| ACC 1h | 109.275 | 36.7243 | 1.13329 | 0.490241 |
| BABA 1h | 84.5338 | 25.4412 | 0.99397 | 0.267839 |
| Chitin 1h | 60.2098 | 5.18428 | 0.844641 | 0.0658528 |
| Epi 1h | 71.3231 | 5.79494 | 1.04073 | 0.103808 |
| SA 1h | 56.8717 | 12.3297 | 0.672398 | 0.102874 |
| Me-JA 1h | 21.569 | 8.31932 | 0.263227 | 0.148925 |
| Control 6h | 74.0725 | 30.6769 | 0.802834 | 0.305183 |
| ABA 6h | 207.754 | 56.9655 | 2.26582 | 0.685779 |
| ACC 6h | 96.0416 | 9.79921 | 1.05376 | 0.0735912 |
| BABA 6h | 62.7849 | 21.324 | 0.641222 | 0.220826 |
| Chitin 6h | 111.024 | 20.4686 | 1.27995 | 0.270475 |
| Epi 6h | 49.7824 | 5.7427 | 0.553676 | 0.0543426 |
| SA 6h | 70.986 | 6.14289 | 0.905652 | 0.0758338 |
| Me-JA 6h | 113.279 | 28.6628 | 1.35525 | 0.326423 |
Source Transcript PGSC0003DMT400004192 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G32450.1 | +3 | 3e-152 | 446 | 217/313 (69%) | nitrate transporter 1.5 | chr1:11715337-11719807 REVERSE LENGTH=614 |
