Probe CUST_27960_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_27960_PI426222305 | JHI_St_60k_v1 | DMT400081889 | GAGCACTTCGAAATGATGTTGTATTCGTGTCAGATCCTCCAACATACATTTAAAAGTATT |
All Microarray Probes Designed to Gene DMG400032148
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_27960_PI426222305 | JHI_St_60k_v1 | DMT400081889 | GAGCACTTCGAAATGATGTTGTATTCGTGTCAGATCCTCCAACATACATTTAAAAGTATT |
| CUST_27958_PI426222305 | JHI_St_60k_v1 | DMT400081888 | GGCACATAATATGCCATGGGATAATGCAGTTAAGGCTGTTTCAAATAATTCTAAGGCAAT |
Microarray Signals from CUST_27960_PI426222305
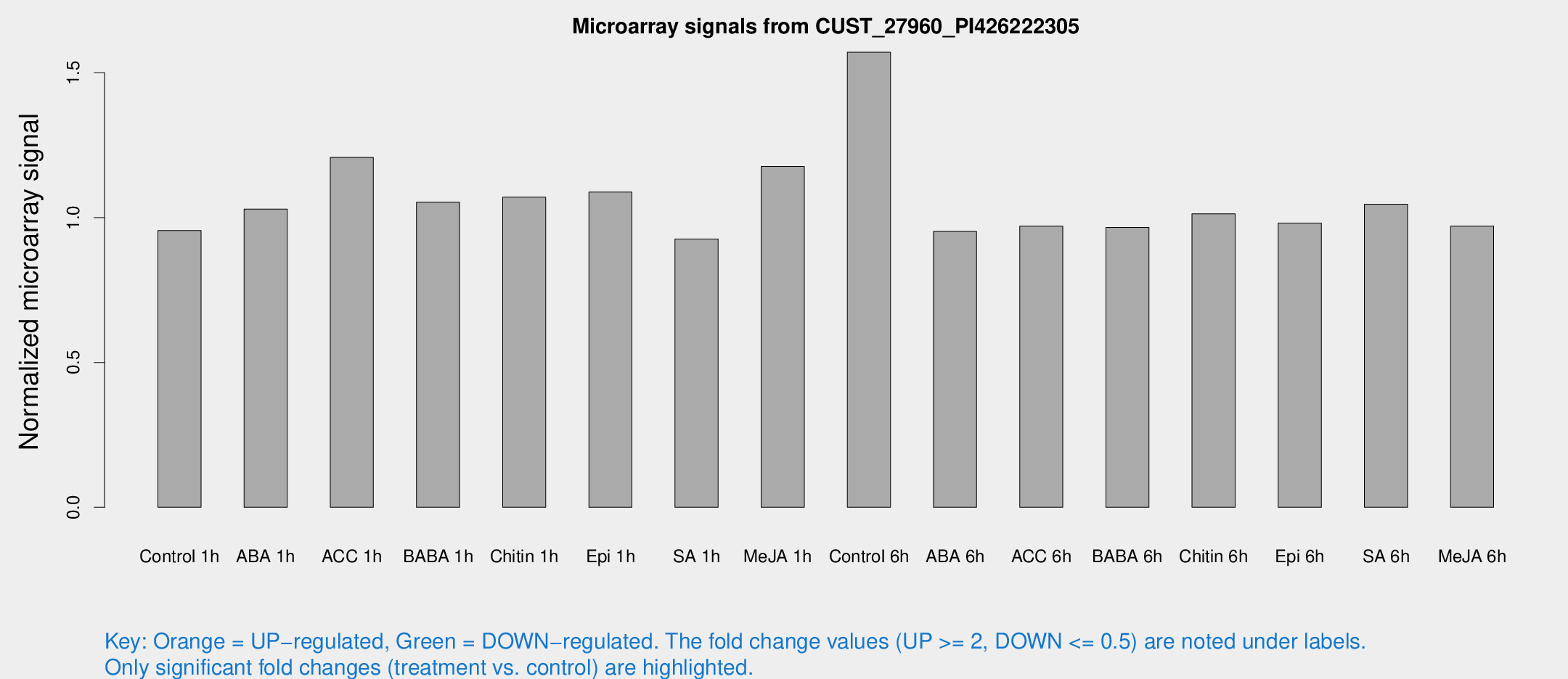
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5.48279 | 3.18052 | 0.955188 | 0.553104 |
| ABA 1h | 5.21914 | 3.02821 | 1.02942 | 0.597124 |
| ACC 1h | 7.46368 | 3.75706 | 1.20767 | 0.633289 |
| BABA 1h | 5.82961 | 3.38794 | 1.0532 | 0.61073 |
| Chitin 1h | 5.42975 | 3.16375 | 1.07032 | 0.606775 |
| Epi 1h | 5.33713 | 3.09871 | 1.088 | 0.621774 |
| SA 1h | 5.45262 | 3.16401 | 0.92578 | 0.536862 |
| Me-JA 1h | 5.53511 | 3.22512 | 1.17635 | 0.681183 |
| Control 6h | 11.6837 | 6.14577 | 1.5707 | 1.21258 |
| ABA 6h | 5.80418 | 3.3683 | 0.951975 | 0.552 |
| ACC 6h | 6.53399 | 3.87958 | 0.970706 | 0.562107 |
| BABA 6h | 6.20809 | 3.60157 | 0.966211 | 0.559514 |
| Chitin 6h | 6.18358 | 3.58825 | 1.01304 | 0.587377 |
| Epi 6h | 6.41411 | 3.76222 | 0.981266 | 0.568503 |
| SA 6h | 5.92981 | 3.43529 | 1.04622 | 0.606045 |
| Me-JA 6h | 5.53913 | 3.21196 | 0.970699 | 0.562812 |
Source Transcript PGSC0003DMT400081889 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G19890.1 | +2 | 7e-99 | 303 | 164/258 (64%) | Peroxidase superfamily protein | chr5:6724372-6725877 REVERSE LENGTH=328 |
