Probe CUST_27330_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_27330_PI426222305 | JHI_St_60k_v1 | DMT400048424 | GCACAATTCATTTCTTGAGTGAATATACATGCTTGCTACTTCAGAACAACAACGATTTGA |
All Microarray Probes Designed to Gene DMG401018810
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_27330_PI426222305 | JHI_St_60k_v1 | DMT400048424 | GCACAATTCATTTCTTGAGTGAATATACATGCTTGCTACTTCAGAACAACAACGATTTGA |
| CUST_27305_PI426222305 | JHI_St_60k_v1 | DMT400048422 | TATATGGAGGCGCAAGAAAGAAATGGGGAAAGAAGGTTTAATGGTCGCCAAGGAGCTTAA |
Microarray Signals from CUST_27330_PI426222305
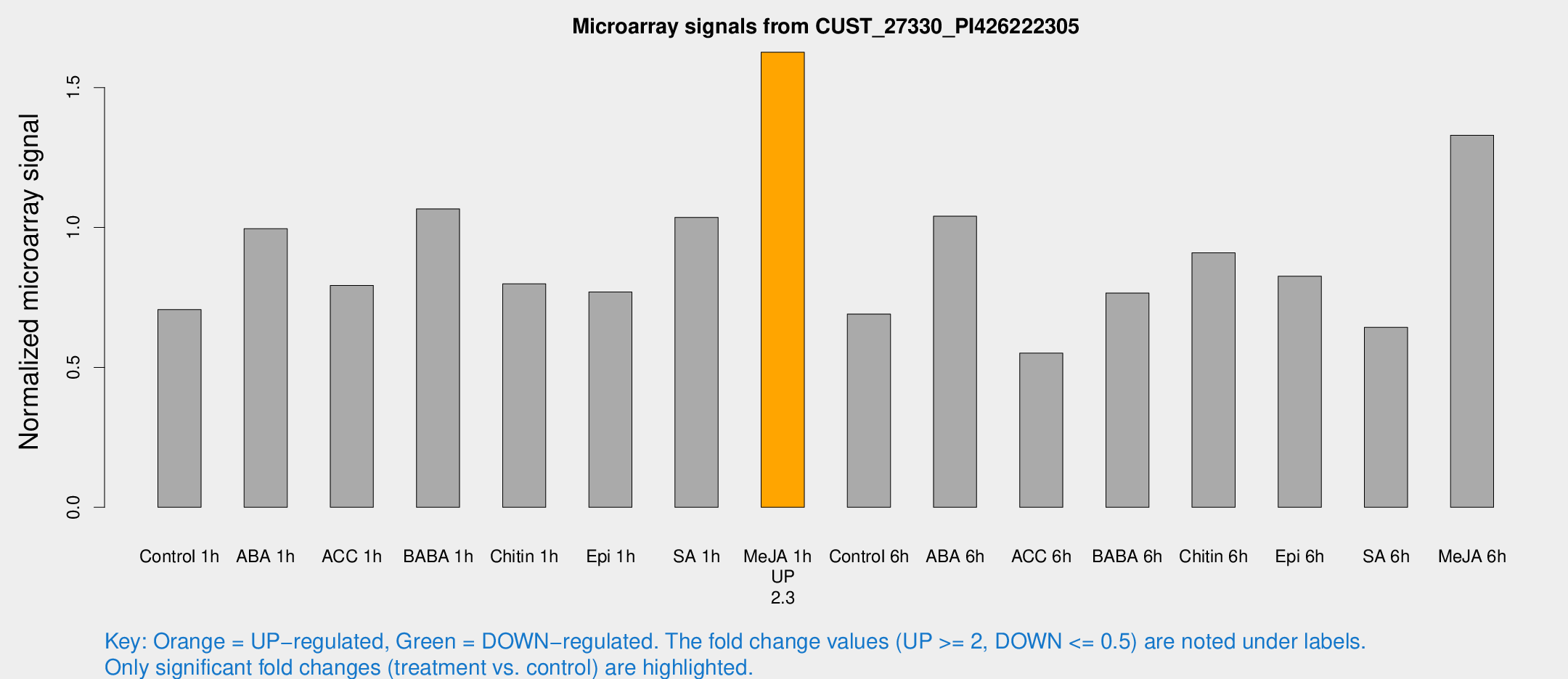
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 44.6035 | 11.9905 | 0.706476 | 0.137344 |
| ABA 1h | 55.7233 | 13.5726 | 0.995622 | 0.241409 |
| ACC 1h | 59.5495 | 22.2566 | 0.792876 | 0.398289 |
| BABA 1h | 61.8811 | 8.39494 | 1.06655 | 0.0908431 |
| Chitin 1h | 43.6475 | 7.54008 | 0.798667 | 0.150277 |
| Epi 1h | 39.5263 | 4.22617 | 0.76962 | 0.0827146 |
| SA 1h | 63.5688 | 6.63773 | 1.03585 | 0.117737 |
| Me-JA 1h | 78.566 | 5.89327 | 1.62616 | 0.121855 |
| Control 6h | 57.0743 | 24.2648 | 0.690668 | 0.509781 |
| ABA 6h | 69.472 | 17.3122 | 1.04007 | 0.202135 |
| ACC 6h | 40.5562 | 11.2148 | 0.550891 | 0.17441 |
| BABA 6h | 57.4771 | 17.0171 | 0.765566 | 0.291515 |
| Chitin 6h | 58.5823 | 8.50737 | 0.909614 | 0.153207 |
| Epi 6h | 58.5781 | 13.0396 | 0.82553 | 0.295137 |
| SA 6h | 40.7269 | 11.1712 | 0.643228 | 0.286543 |
| Me-JA 6h | 81.7654 | 16.3717 | 1.3296 | 0.171102 |
Source Transcript PGSC0003DMT400048424 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G62350.1 | +1 | 3e-68 | 230 | 110/140 (79%) | Pentatricopeptide repeat (PPR) superfamily protein | chr1:23056653-23057935 FORWARD LENGTH=196 |
