Probe CUST_27233_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_27233_PI426222305 | JHI_St_60k_v1 | DMT400013766 | ATCTTTGCAAAATCAAGTTGAGTTCCTCTCAATGAAGCTATCTGCAGCTAGCACATATTA |
All Microarray Probes Designed to Gene DMG400005388
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_27213_PI426222305 | JHI_St_60k_v1 | DMT400013765 | GTGAGTTGAAATGATCCAACACAACTTCTTTTCTGATTGACCAAAACACACACACAAAAA |
| CUST_27233_PI426222305 | JHI_St_60k_v1 | DMT400013766 | ATCTTTGCAAAATCAAGTTGAGTTCCTCTCAATGAAGCTATCTGCAGCTAGCACATATTA |
Microarray Signals from CUST_27233_PI426222305
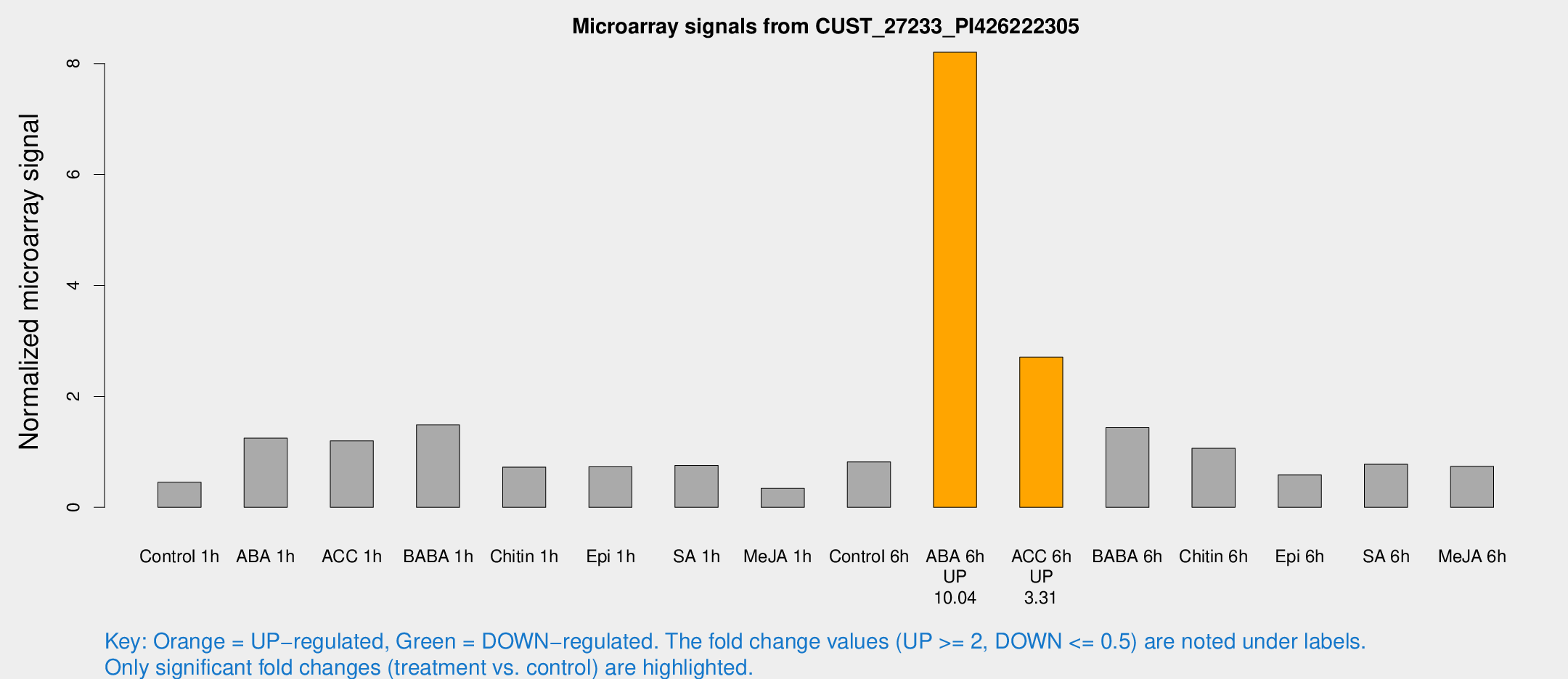
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 14.3662 | 6.55677 | 0.450717 | 0.200047 |
| ABA 1h | 42.1706 | 22.4667 | 1.24638 | 1.14738 |
| ACC 1h | 36.2915 | 11.084 | 1.1983 | 0.41146 |
| BABA 1h | 38.5893 | 7.40481 | 1.48519 | 0.184033 |
| Chitin 1h | 21.308 | 10.4079 | 0.725273 | 0.362185 |
| Epi 1h | 19.729 | 6.96682 | 0.729846 | 0.408491 |
| SA 1h | 22.2595 | 7.6761 | 0.754916 | 0.210604 |
| Me-JA 1h | 7.42423 | 3.68529 | 0.340602 | 0.175624 |
| Control 6h | 23.4731 | 7.46095 | 0.817573 | 0.224247 |
| ABA 6h | 231.633 | 36.8301 | 8.20547 | 1.09636 |
| ACC 6h | 87.3064 | 22.3658 | 2.70728 | 0.748926 |
| BABA 6h | 46.744 | 14.9651 | 1.43672 | 0.495708 |
| Chitin 6h | 29.7157 | 4.4314 | 1.06543 | 0.163348 |
| Epi 6h | 19.185 | 7.06636 | 0.583606 | 0.219726 |
| SA 6h | 23.2585 | 8.20408 | 0.774603 | 0.267552 |
| Me-JA 6h | 20.5881 | 6.22406 | 0.736488 | 0.242662 |
Source Transcript PGSC0003DMT400013766 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G18400.1 | +3 | 2e-56 | 187 | 120/257 (47%) | BR enhanced expression 1 | chr1:6331464-6333576 FORWARD LENGTH=260 |
