Probe CUST_26817_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_26817_PI426222305 | JHI_St_60k_v1 | DMT400093941 | AGAAAATCATCGTTAATCCCATGCCAATTCCGGCTAACGGACTTCCGATCAGACTCTATC |
All Microarray Probes Designed to Gene DMG400043512
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_26817_PI426222305 | JHI_St_60k_v1 | DMT400093941 | AGAAAATCATCGTTAATCCCATGCCAATTCCGGCTAACGGACTTCCGATCAGACTCTATC |
Microarray Signals from CUST_26817_PI426222305
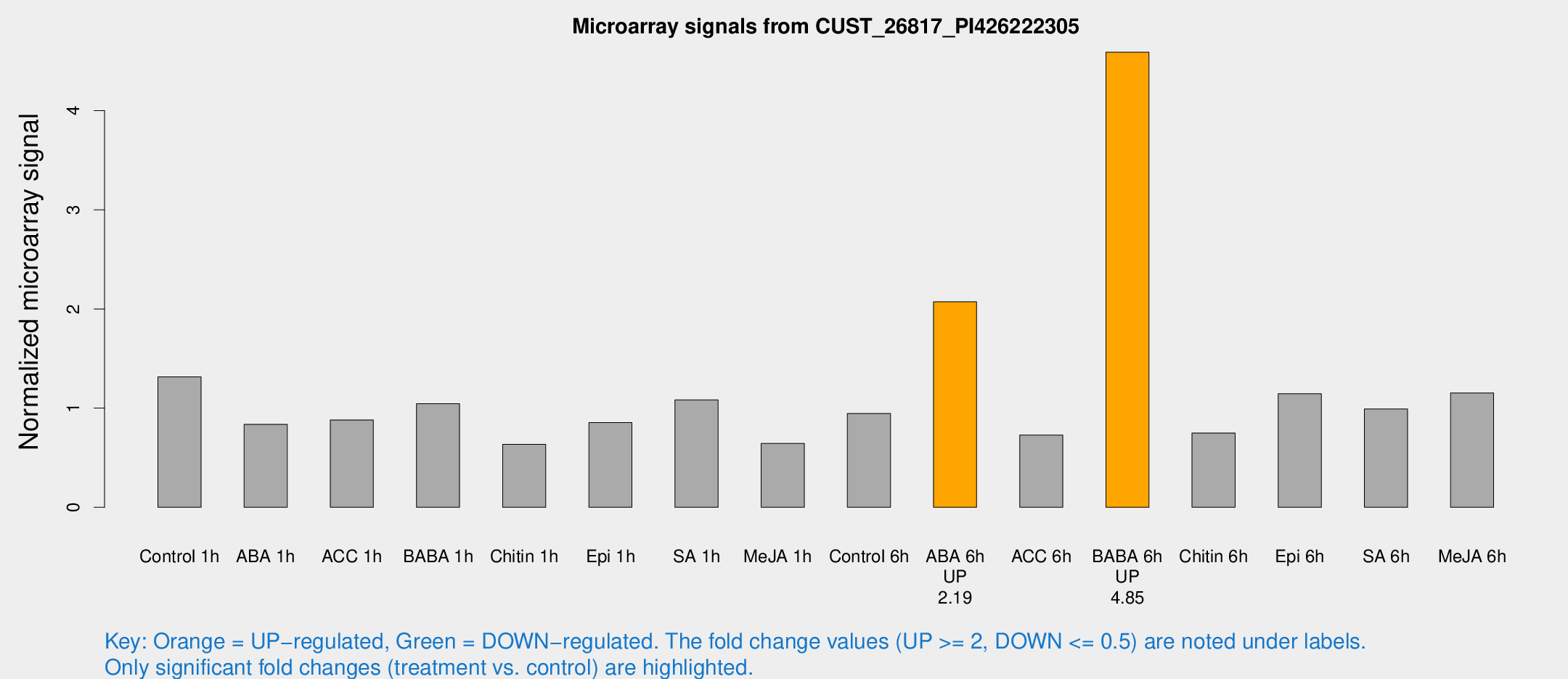
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 15.0772 | 3.13748 | 1.31609 | 0.30747 |
| ABA 1h | 9.41344 | 3.83169 | 0.836749 | 0.367075 |
| ACC 1h | 11.7979 | 5.15359 | 0.880622 | 0.356054 |
| BABA 1h | 12.4373 | 3.85857 | 1.04526 | 0.373434 |
| Chitin 1h | 6.342 | 3.07223 | 0.635398 | 0.32056 |
| Epi 1h | 8.33163 | 3.081 | 0.853952 | 0.34562 |
| SA 1h | 12.4181 | 3.12153 | 1.0836 | 0.284872 |
| Me-JA 1h | 5.77972 | 3.12347 | 0.643364 | 0.350036 |
| Control 6h | 11.3057 | 3.24207 | 0.945335 | 0.342629 |
| ABA 6h | 25.2605 | 5.38213 | 2.07202 | 0.345098 |
| ACC 6h | 12.2258 | 6.77871 | 0.728711 | 0.340589 |
| BABA 6h | 56.676 | 5.05811 | 4.58851 | 0.394571 |
| Chitin 6h | 9.41747 | 3.52293 | 0.748732 | 0.327663 |
| Epi 6h | 15.3077 | 4.49482 | 1.14566 | 0.331936 |
| SA 6h | 11.6923 | 3.41144 | 0.990843 | 0.435483 |
| Me-JA 6h | 14.793 | 4.81896 | 1.1534 | 0.469833 |
Source Transcript PGSC0003DMT400093941 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT5G36110.1 | +1 | 9e-156 | 450 | 211/346 (61%) | cytochrome P450, family 716, subfamily A, polypeptide 1 | chr5:14195377-14197613 FORWARD LENGTH=477 |
