Probe CUST_26531_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_26531_PI426222305 | JHI_St_60k_v1 | DMT400001025 | TGGGGAACAGCCAATAACAGTTTCATCGTTGTTGAACCTTTGGATTTCTCTCAACGAATT |
All Microarray Probes Designed to Gene DMG400000380
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_26531_PI426222305 | JHI_St_60k_v1 | DMT400001025 | TGGGGAACAGCCAATAACAGTTTCATCGTTGTTGAACCTTTGGATTTCTCTCAACGAATT |
Microarray Signals from CUST_26531_PI426222305
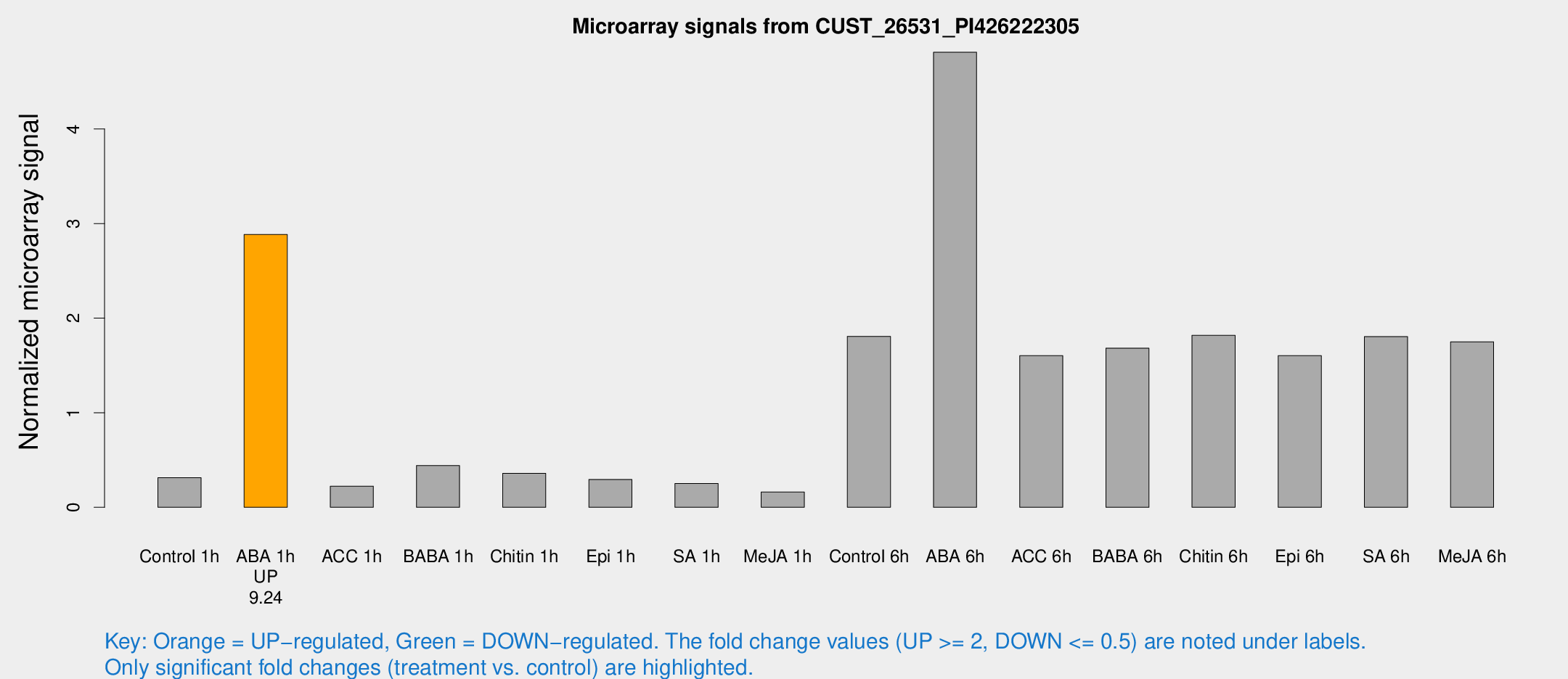
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 50.9873 | 6.5419 | 0.312329 | 0.0273546 |
| ABA 1h | 431.21 | 90.4013 | 2.88549 | 0.574903 |
| ACC 1h | 38.5658 | 7.64264 | 0.223259 | 0.035889 |
| BABA 1h | 71.4465 | 14.4115 | 0.441509 | 0.054231 |
| Chitin 1h | 52.7016 | 7.11933 | 0.358693 | 0.0447207 |
| Epi 1h | 41.3893 | 4.61368 | 0.294451 | 0.0302654 |
| SA 1h | 42.6337 | 6.791 | 0.251385 | 0.0330358 |
| Me-JA 1h | 22.5506 | 5.8128 | 0.16146 | 0.0302992 |
| Control 6h | 335.296 | 118.425 | 1.80801 | 0.615473 |
| ABA 6h | 889.668 | 220.99 | 4.81208 | 1.30909 |
| ACC 6h | 316.252 | 73.841 | 1.60323 | 0.359657 |
| BABA 6h | 330.91 | 90.7182 | 1.68265 | 0.507849 |
| Chitin 6h | 322.992 | 60.1217 | 1.81816 | 0.402805 |
| Epi 6h | 302.743 | 60.1117 | 1.60305 | 0.446264 |
| SA 6h | 324.439 | 97.5132 | 1.80452 | 0.934921 |
| Me-JA 6h | 313.671 | 93.4567 | 1.74959 | 0.516641 |
Source Transcript PGSC0003DMT400001025 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT3G24520.1 | +2 | 7e-86 | 271 | 133/203 (66%) | heat shock transcription factor C1 | chr3:8941455-8942531 FORWARD LENGTH=330 |
