Probe CUST_26271_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_26271_PI426222305 | JHI_St_60k_v1 | DMT400041686 | GCAGAGTCCTTGGTATCTGAAAAGAAGTTTTGGCAAGCTTGTATCATTTTTTGTTCTAAA |
All Microarray Probes Designed to Gene DMG400016162
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_26271_PI426222305 | JHI_St_60k_v1 | DMT400041686 | GCAGAGTCCTTGGTATCTGAAAAGAAGTTTTGGCAAGCTTGTATCATTTTTTGTTCTAAA |
| CUST_26200_PI426222305 | JHI_St_60k_v1 | DMT400041687 | TGTGTAGTTCGAAGTAGTAGCACTTTGATCTTCATCATATGGAAATTTAACGCCTATTTG |
Microarray Signals from CUST_26271_PI426222305
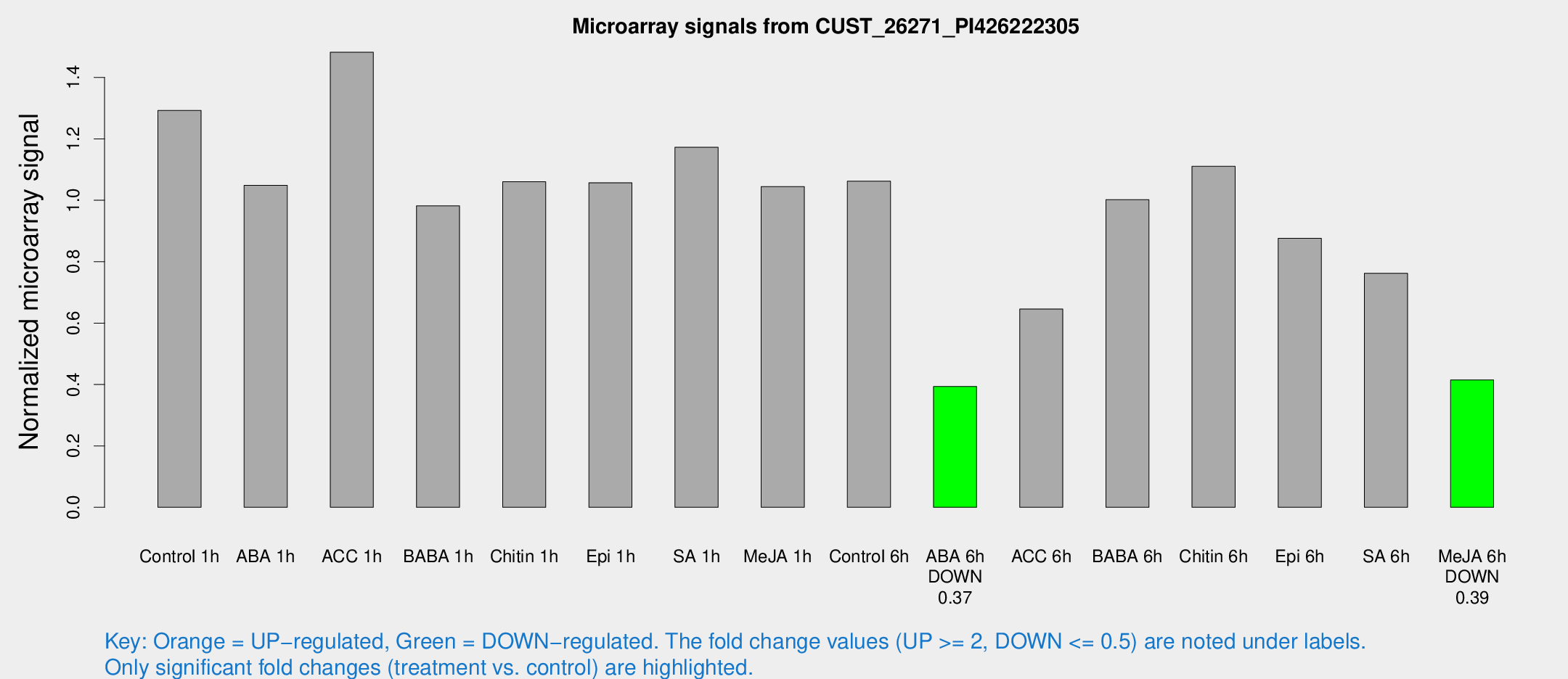
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 5830.09 | 708.55 | 1.29284 | 0.0755726 |
| ABA 1h | 4254.81 | 733.633 | 1.04854 | 0.23226 |
| ACC 1h | 6787.61 | 418.734 | 1.48223 | 0.165148 |
| BABA 1h | 4253.31 | 446.399 | 0.981866 | 0.0566959 |
| Chitin 1h | 4307.63 | 582.866 | 1.06029 | 0.163003 |
| Epi 1h | 4072.77 | 235.732 | 1.05696 | 0.0610312 |
| SA 1h | 5368.71 | 317.237 | 1.17259 | 0.119925 |
| Me-JA 1h | 3797.37 | 219.767 | 1.0448 | 0.061338 |
| Control 6h | 5011.89 | 1090.17 | 1.06239 | 0.175032 |
| ABA 6h | 1863.91 | 108.009 | 0.39352 | 0.0253738 |
| ACC 6h | 3461.46 | 764.333 | 0.645837 | 0.0741981 |
| BABA 6h | 5187.67 | 1029.41 | 1.00215 | 0.227032 |
| Chitin 6h | 5335.83 | 696.261 | 1.1105 | 0.106061 |
| Epi 6h | 4411.05 | 321.364 | 0.876319 | 0.12744 |
| SA 6h | 3489.91 | 660.841 | 0.762297 | 0.0725441 |
| Me-JA 6h | 1994.1 | 519.346 | 0.414707 | 0.0945769 |
Source Transcript PGSC0003DMT400041686 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT4G39970.1 | +2 | 7e-151 | 434 | 208/255 (82%) | Haloacid dehalogenase-like hydrolase (HAD) superfamily protein | chr4:18536678-18538429 REVERSE LENGTH=316 |
