Probe CUST_26183_PI426222305 - General Information
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_26183_PI426222305 | JHI_St_60k_v1 | DMT400041725 | ATTATATAGCAGGAGCAATCTCGATTGCCGTACCAGAGAGGAGAGGAATATTATATAACA |
All Microarray Probes Designed to Gene DMG400016179
| Probe ID | Chip name | Transcript ID | Probe Sequence |
|---|---|---|---|
| CUST_26183_PI426222305 | JHI_St_60k_v1 | DMT400041725 | ATTATATAGCAGGAGCAATCTCGATTGCCGTACCAGAGAGGAGAGGAATATTATATAACA |
Microarray Signals from CUST_26183_PI426222305
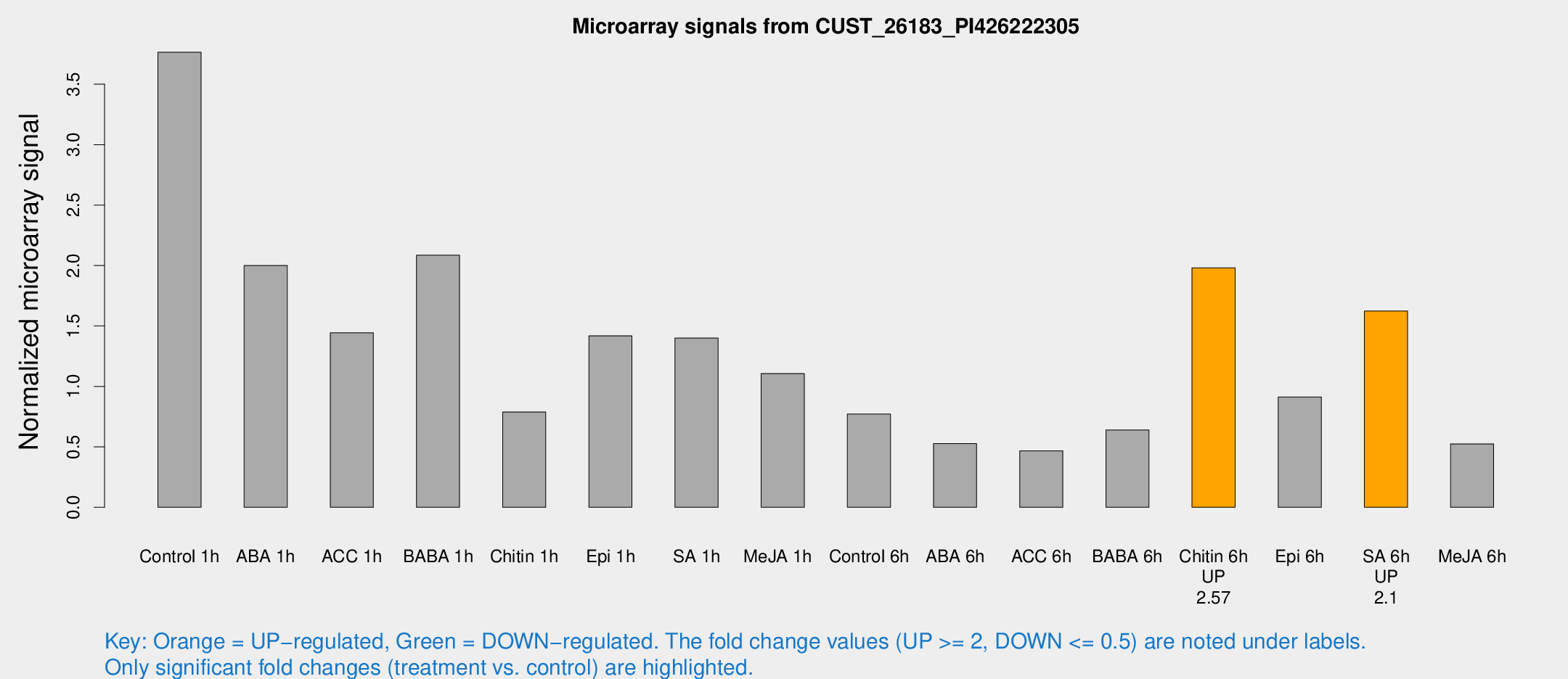
| Treatment | Raw signal | Raw Std Err | Normalized signal | Normalized Std Err |
|---|---|---|---|---|
| Control 1h | 325.581 | 73.3679 | 3.76457 | 0.574656 |
| ABA 1h | 213.696 | 107.75 | 1.99978 | 1.70209 |
| ACC 1h | 170.861 | 77.7359 | 1.44421 | 1.07809 |
| BABA 1h | 186.966 | 56.8921 | 2.08572 | 0.584604 |
| Chitin 1h | 61.6492 | 14.7977 | 0.788414 | 0.13728 |
| Epi 1h | 111.492 | 31.4213 | 1.41831 | 0.463101 |
| SA 1h | 121.417 | 17.485 | 1.40002 | 0.30368 |
| Me-JA 1h | 85.3701 | 33.121 | 1.10657 | 0.336946 |
| Control 6h | 64.2621 | 5.72794 | 0.771339 | 0.0605003 |
| ABA 6h | 47.0974 | 6.10028 | 0.52721 | 0.104679 |
| ACC 6h | 45.3867 | 7.24493 | 0.467604 | 0.138453 |
| BABA 6h | 83.8398 | 50.0193 | 0.640104 | 0.462922 |
| Chitin 6h | 174.317 | 10.7282 | 1.98121 | 0.130229 |
| Epi 6h | 92.9644 | 29.342 | 0.912697 | 0.363515 |
| SA 6h | 133.053 | 8.76495 | 1.62352 | 0.274577 |
| Me-JA 6h | 50.9399 | 17.9275 | 0.524797 | 0.205406 |
Source Transcript PGSC0003DMT400041725 - Homology to Model Species (BLASTX to E-value < 1e-50)
| Database | Link to BLAST Hit | Frame | E-value | Score | % Identity | Description |
|---|---|---|---|---|---|---|
| Tomato (ITAG) | None | - | - | - | - | - |
| TAIR PP10 | AT1G65480.1 | +2 | 1e-93 | 275 | 126/173 (73%) | PEBP (phosphatidylethanolamine-binding protein) family protein | chr1:24331510-24333689 FORWARD LENGTH=175 |
